
Sample to Insight__
691223
QIAGEN GmbH, QIAGEN Strasse 1, D
R
September 2020
QIAstat-Dx® Respiratory
SARS-CoV-2 Panel
Instructions for Use
(Handbook)
6
Version 1
For
in vitro
Rx Only
4
diagnostic use under Emergency Use Authorization Only
-40724 Hilden

2
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Contents
Intended Use .............................................................................................................. 4
Summary and Explanation ........................................................................................... 6
QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge Description ............................ 6
Pathogen Information .................................................................................................. 8
Principle of the Procedure .......................................................................................... 10
Description of the process ................................................................................ 10
Sample collection and cartridge loading ........................................................... 11
Sample preparation, nucleic acid amplification and detection ............................. 13
Materials Provided .................................................................................................... 14
Kit contents .................................................................................................... 14
Materials Required but Not Provided........................................................................... 15
Warnings and Precautions ......................................................................................... 16
Safety information .......................................................................................... 16
Reagent Storage and Handling .................................................................................. 19
Specimen Handling, Storage and Preparation .............................................................. 19
Procedure ................................................................................................................ 20
Internal Control .............................................................................................. 20
Protocol: Transport medium liquid samples ........................................................ 21
Viewing results ............................................................................................... 32
Interpretation of Results .............................................................................................. 34
Internal Control interpretation ........................................................................... 34
Pathogen Result interpretation .......................................................................... 34

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
3
Quality Control ......................................................................................................... 43
Limitations ................................................................................................................ 44
Conditions of Authorization for the Laboratory ............................................................. 47
Performance Characteristics ....................................................................................... 49
SARS-CoV-2 Target ......................................................................................... 49
Clinical performance ...................................................................................... 49
Analytical performance ................................................................................... 50
Additional Targets included in the QIAstat-Dx SARS-CoV-2 Panel ......................... 53
Clinical performance ...................................................................................... 53
Expected values ............................................................................................. 73
Analytical performance ................................................................................... 80
Appendices............................................................................................................ 111
Appendix A: Installing the Assay Definition File ................................................ 111
Appendix B: Glossary ................................................................................... 113
Appendix C: Disclaimer of warranties ............................................................. 115
Symbols ................................................................................................................. 116
Ordering Information .............................................................................................. 117
Handbook Revision History ...................................................................................... 119

4
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Intended Use
The QIAstat-Dx Respiratory SARS-CoV-2 Panel is a multiplexed nucleic acid real-time PCR test
intended for the qualitative detection and differentiation of nucleic acid from multiple
respiratory viral and bacterial organisms, including the SARS-CoV-2 virus, in nasopharyngeal
swabs (NPS) eluted in universal transport media collected from patients suspected of
COVID-19 by their healthcare provider. Testing is limited to laboratories certified under the
Clinical Laboratory Improvement Amendments of 1988 (CLIA), 42 U.S.C. §263a, to perform
high complexity and moderate complexity tests.
The QIAstat-Dx Respiratory SARS-CoV-2 Panel is intended for the detection and differentiation
of nucleic acid from SARS-CoV-2 and the following organism types and subtypes: Adenovirus,
Coronavirus 229E, Coronavirus HKU1, Coronavirus NL63, Coronavirus OC43, SARS-CoV-2,
Human Metapneumovirus A+B, Influenza A, Influenza A H1, Influenza A H3, Influenza A
H1N1/pdm09, Influenza B, Parainfluenza virus 1, Parainfluenza virus 2, Parainfluenza virus
3, Parainfluenza virus 4, Rhinovirus/Enterovirus, Respiratory Syncytial Virus A+B,
,
pertussis
Chlamydophila pneumoniae
, and
Mycoplasma pneumoniae
.
Bordetella
SARS-CoV-2 RNA and nucleic acids from the other respiratory viral and bacterial organisms
identified by this test are generally detectable in nasopharyngeal swabs (NPS) during the acute
phase of infection. Positive results are indicative of the presence of the identified
microorganism, but do not rule out co-infection with other pathogens not detected by the test,
or lower respiratory tract infection that is not detected by a nasopharyngeal swab. The agent
detected may not be the definite cause of disease.
For SARS-CoV-2 positive specimens; clinical correlation with patient history and other
diagnostic information is necessary to determine patient infection status. Laboratories within
the United States and its territories are required to report all SARS-CoV-2 positive results to the
appropriate public health authorities.

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
5
Negative results do not preclude SARS-CoV-2 infection and should not be used as the sole
basis for treatment or other patient management decisions. Negative SARS-CoV-2 results must
be combined with clinical observations, patient history, and epidemiological information.
Negative results for other organisms identified by the test may require additional laboratory
testing (eg, bacterial and viral culture, immunofluorescence and radiography) when evaluating
a patient with possible respiratory tract infection.
Testing with the QIAstat Dx Respiratory SARS-CoV-2 Panel is intended for use by qualified and
trained operators who are proficient in performing the tests using the QIAstat Dx Analyzer
System. The QIAstat Dx Respiratory SARS-CoV-2 Panel is only for use under the Food and Drug
Administration’s Emergency Use Authorization.

6
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Summary and Explanation
QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge Description
The QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge is a disposable plastic device that
allows performance of fully automated molecular assays for the detection of respiratory
pathogens. The main features of the QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge
include compatibility with nasopharyngeal swab in transport medium (liquid samples),
hermetical containment of the pre-loaded reagents necessary for testing, and true walk-away
operation. All sample preparation and assay testing steps are performed within the cartridge.
All reagents required for the complete execution of a test run are pre-loaded and self-contained in
the QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge. The user does not need to come in contact
with and/or manipulate any reagents. During the test, reagents are handled within the cartridge in
the Analytical Module of the QIAstat-Dx Analyzer 1.0 by pneumatically-operated microfluidics and
make no direct contact with the actuators. The QIAstat-Dx Analyzer 1.0 houses air filters for both
incoming and outgoing air, further safeguarding the environment. After testing, the cartridge stays
hermetically closed at all times, greatly enhancing its safe disposal.
Within the cartridge, multiple steps are automatically performed in sequence using pneumatic
pressure to transfer samples and fluids via the transfer chamber to their intended destinations.

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
7
Swab port
Main port
Traceability bar code
Reaction chambers
After the QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge containing the sample is
introduced into the QIAstat-Dx Analyzer 1.0, the following assay steps occur automatically:
Resuspension of Internal Control
Cell lysis using mechanical and/or chemical means
Membrane-based nucleic acid purification
Mixing of the purified nucleic acid with lyophilized master mix reagents
Transfer of defined aliquots of eluate/master mix to different reaction chambers
Performance of multiplex real-time RT-PCR testing within each reaction chamber.
Note: An increase in fluorescence, indicating detection of the target analyte, is detected
directly within each reaction chamber.
Figure 1. Layout of the QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge and its features.
Note: The swab port is not used for the QIAstat-Dx Respiratory SARS-CoV-2 Panel assay.

8
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Pathogen Information
Acute respiratory infections can be caused by a variety of pathogens, including bacteria and
viruses, and generally present with nearly indistinguishable clinical signs and symptoms. The
rapid and accurate determination of the presence or absence of potential causative agent(s)
helps make timely decisions regarding treatment, hospital admission, infection control, and
return of the patient to work and family. It may also greatly support improved antimicrobial
stewardship and other important public health initiatives.
The QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge is a single-use cartridge that includes
all reagents needed for nucleic acid extraction, nucleic acid amplification and detection of
22 bacteria and viruses (or their subtypes), including SARS-CoV-2 that cause respiratory
symptoms. Testing requires a small sample volume and minimal hands-on time, and the results
are available in approximately one hour.
The SARS-CoV-2 target in the QIAstat-Dx Respiratory SARS-CoV-2 Panel has been designed
upon alignment of more than 170 genomic sequences available in public databases from the
SARS-CoV-2, which was identified as the causative agent of the viral pneumonia (COVID-19)
outbreak originated in Wuhan, Hubei, China. The SARS-CoV-2 in this panel targets 2 genes
of the virus genome (Orf1b poly gene (Rdrp gene) and E genes) detected with the same
fluorescence channel. The two gene targets are not differentiated and amplification of either
or both gene targets leads to a fluorescence signal.
Pathogens (and subtypes) that can be detected and identified with the QIAstat-Dx Respiratory
SARS-CoV-2 Panel are listed in Table 1 (next page).

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
9
Table 1. Pathogens detected by the QIAstat-Dx Respiratory SARS-CoV-2 Panel
Pathogen Classification (genome type)
Influenza A Orthomyxovirus (RNA)
Influenza A, subtype H1N1/2009/pdm09 Orthomyxovirus (RNA)
Influenza A subtype H1 Orthomyxovirus (RNA)
Influenza A subtype H3 Orthomyxovirus (RNA)
Influenza B Orthomyxovirus (RNA)
Coronavirus 229E Coronavirus (RNA)
Coronavirus HKU1 Coronavirus (RNA)
Coronavirus NL63 Coronavirus (RNA)
Coronavirus OC43 Coronavirus (RNA)
SARS-CoV-2 Coronavirus (RNA)
Parainfluenza virus 1 Paramyxovirus (RNA)
Parainfluenza virus 2 Paramyxovirus (RNA)
Parainfluenza virus 3 Paramyxovirus (RNA)
Parainfluenza virus 4 Paramyxovirus (RNA)
Respiratory Syncytial Virus A/B Paramyxovirus (RNA)
Human Metapneumovirus A/B Paramyxovirus (RNA)
Adenovirus Adenovirus (DNA)
Rhinovirus/Enterovirus Picornavirus (RNA)
Mycoplasma pneumoniae
Chlamydophila pneumoniae
Bordetella pertussis
Bacterium (DNA)
Bacterium (DNA)
Bacterium (DNA)
Note: Enterovirus and Rhinovirus are both detected, but not differentiated, with the QIAstat-Dx Respiratory SARS-CoV-2
Panel.
10
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Principle of the Procedure
Description of the process
Diagnostic tests with the QIAstat-Dx Respiratory SARS-CoV-2 Panel are performed on the
QIAstat-Dx Analyzer 1.0. All of the sample preparation and analysis steps are performed
automatically by the QIAstat-Dx Analyzer 1.0. Samples are collected and loaded manually
into the QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge:
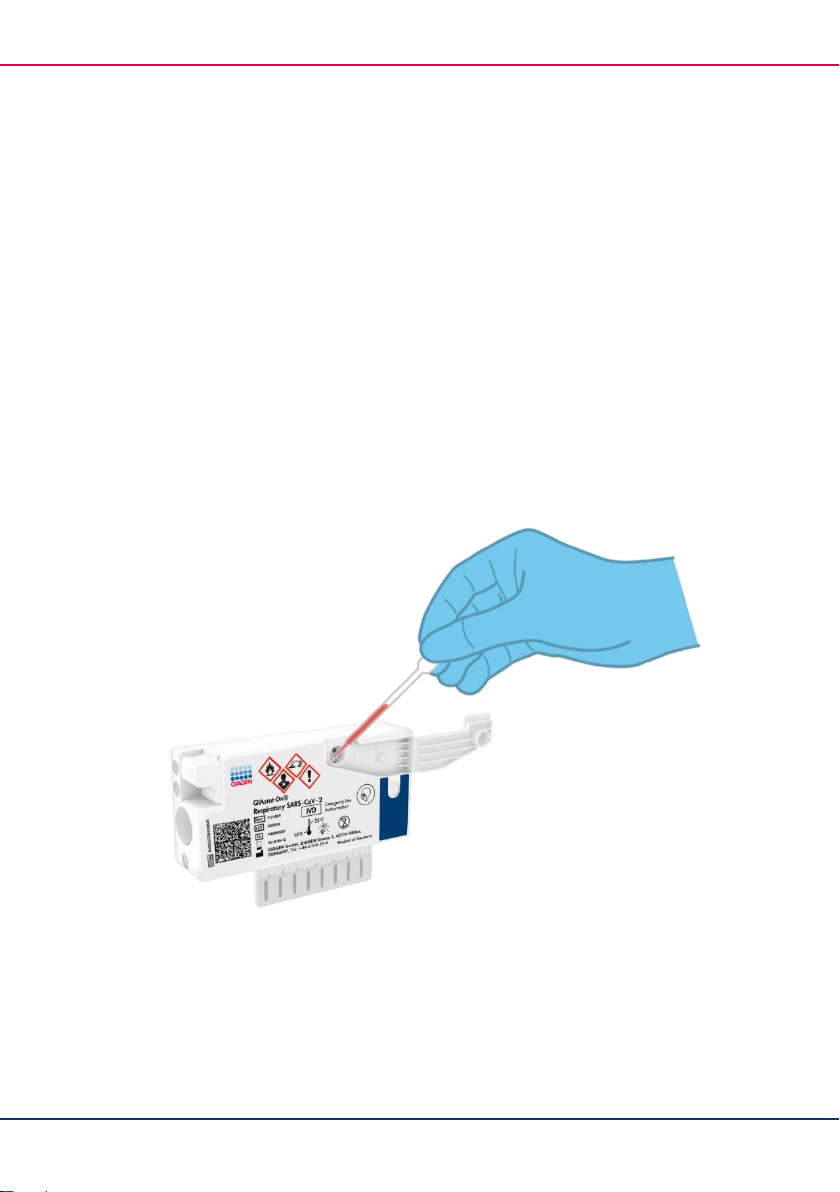
A transfer pipette provided with the test kit is used for dispensing transport medium liquid
sample into the main port (Figure 2).
Figure 2. Dispensing transport medium liquid sample into the main port.

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
11
Sample collection and cartridge loading
The collection of samples and their subsequent loading into the QIAstat-Dx Respiratory
SARS-CoV-2 Panel Cartridge should be performed by personnel trained in safe handling of
biological samples.
The following steps are involved and must be executed by the user:
1. A nasopharyngeal swab sample is collected.
2. The nasopharyngeal swab is placed into transport medium.
3. The sample information is manually written on or a sample label is affixed to the top of a
QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge.
4. Transport medium liquid sample is loaded manually into the QIAstat-Dx Respiratory
SARS-CoV-2 Panel Cartridge.
300 μl of sample is transferred into the main port of the QIAstat-Dx Respiratory
SARS-CoV-2 Panel Cartridge using one of the included transfer pipettes.
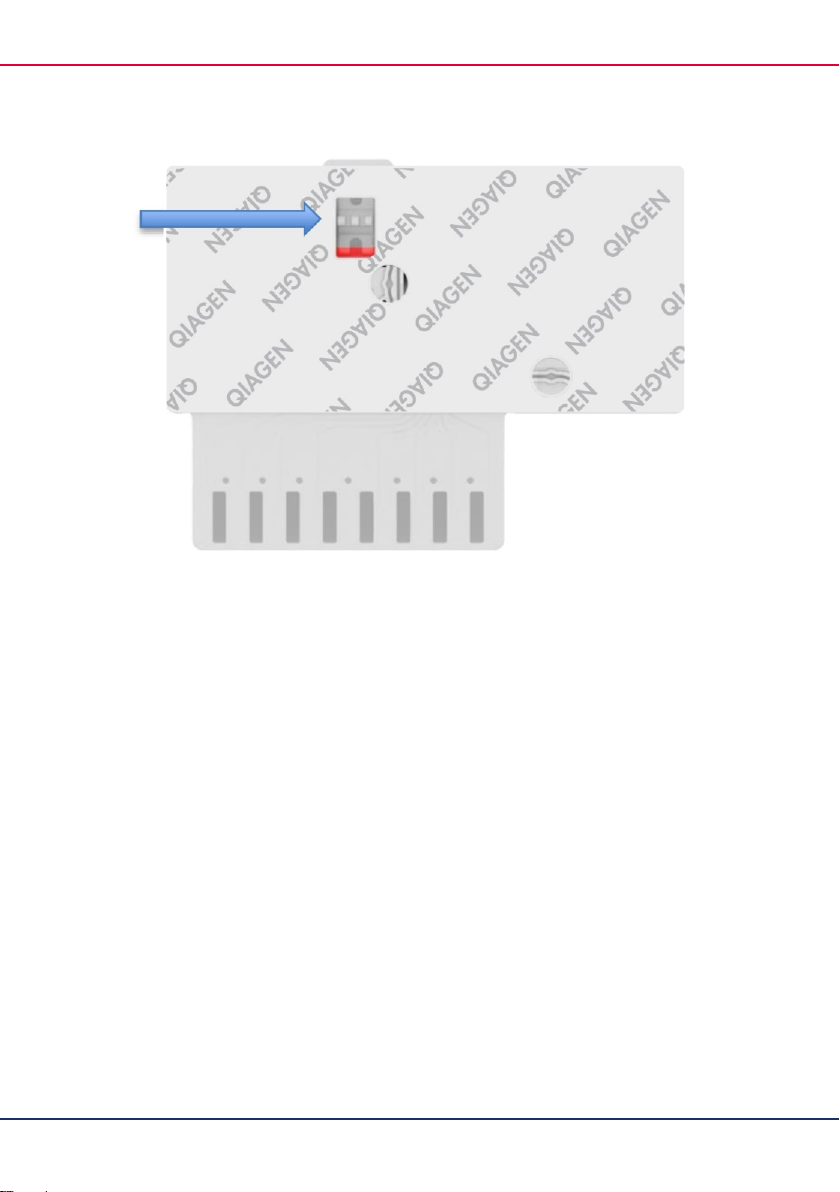
Note: When loading transport medium liquid sample, the user performs a visual check of
the sample inspection window (see image below) to confirm that the liquid sample has
been loaded (Figure 3, next page).
12
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Figure 3. Sample inspection window (blue arrow).
5. The sample bar code and QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge QR code
are scanned in the QIAstat-Dx Analyzer 1.0.
6. The QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge is introduced into the QIAstat-Dx
Analyzer 1.0.
7. The test is started on the QIAstat-Dx Analyzer 1.0.

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
13
Sample preparation, nucleic acid amplification and detection
The extraction, amplification, and detection of nucleic acids in the sample are performed
automatically by the QIAstat-Dx Analyzer 1.0.
1. The liquid sample is homogenized and cells are lysed in the lysis chamber of the QIAstatDx Respiratory SARS-CoV-2 Panel Cartridge, which includes a rotor that turns at high
speed.
2. Nucleic acids are purified from the lysed sample via binding to a silica membrane in the
purification chamber of the QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge in the
presence of chaotropic salts and alcohol.
3. The purified nucleic acids are eluted from the membrane in the purification chamber and
are mixed with the lyophilized PCR chemistry in the dried-chemistry chamber of the
QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge.
4. The mixture of sample and PCR reagents is dispensed into the QIAstat-Dx Respiratory
SARS-CoV-2 Panel Cartridge PCR chambers, which contain lyophilized, assay-specific
primers and probes.
5. The QIAstat-Dx Analyzer 1.0 creates the optimal temperature profiles to carry out
effective multiplex real-time RT-PCR and performs real-time fluorescence measurements to
generate amplification curves.
6. The QIAstat-Dx Analyzer 1.0 Software interprets the resulting data and process controls
and delivers a test report.
14
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Materials Provided
Kit contents
QIAstat-Dx Respiratory SARS-CoV-2 Panel
Catalog no.
Number of tests
QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge* 6
Transfer pipettes† 6
* 6 individually packaged cartridges containing all reagents needed for sample preparation and multiplex real-time RT-
PCR, plus Internal Control.
†
6 individually packaged transfer pipettes for dispensing liquid sample into the QIAstat-Dx Respiratory SARS-CoV-2
Panel Cartridge.
691223
6

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
15
Materials Required but Not Provided
The QIAstat-Dx Respiratory SARS-CoV-2 Panel is designed for use with the QIAstat-Dx Analyzer
1.0. Before beginning a test, make sure the following are available:
QIAstat-Dx Analyzer 1.0 (at least one Operational Module and one Analytical Module)
with software version 1.2 or higher
QIAstat-Dx Analyzer 1.0 User Manual
QIAstat-Dx latest Assay Definition File software for Respiratory SARS-CoV-2 Panel
installed on the Operational Module
(for use with software version 1.2 or higher)

16
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Warnings and Precautions
For
in vitro
diagnostic use under Emergency Use Authorization only.
The QIAstat-Dx Respiratory SARS-CoV-2 Panel is to be used by laboratory professionals trained
in the use of QIAstat-Dx Analyzer 1.0.
This device is restricted to sale by or on the order of a physician, or to a clinical laboratory;
its use is restricted to, by, or on the order of a physician.
Pertussis is a nationally notifiable infectious disease in the U.S. If
detected, notify state and/or local health departments.
Laboratories are required to report all positive SARS-CoV-2 results to the appropriate public
health authorities.
Bordetella pertussis
is
Safety information
When working with chemicals, always wear a suitable lab coat, disposable gloves, and
protective goggles. For more information, consult the appropriate safety data sheets (SDSs).
These are available online in PDF format at www.qiagen.com/safety where you can find, view
and print the SDS for each QIAGEN kit and kit component.
Always wear appropriate personal protective equipment, including but not limited to
disposable powder-free gloves, a lab coat, and protective eyewear. Protect skin, eyes, and
mucus membranes. Change gloves often when handling samples.

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
17
Handle all samples, used cartridges and transfer pipettes as if they are capable of transmitting
infectious agents. Always observe safety precautions as outlined in relevant guidelines, such
®
(CLSI)
as the Clinical and Laboratory Standards Institute
from Occupationally Acquired Infections; Approved Guideline
Protection of Laboratory Workers
(M29), or other appropriate
documents provided by:
OSHA
ACGIH
®
: Occupational Safety and Health Administration (United States of America)
®
: American Conference of Government Industrial Hygienists (United States of
America)
COSHH: Control of Substances Hazardous to Health (United Kingdom)
Follow your institution’s safety procedures for handling biological samples. Dispose of
samples, QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridges, and transfer pipettes
according to the appropriate regulations.
The QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge is a closed, single-use device that contains
all reagents needed for sample preparation and multiplex real-time RT-PCR within the QIAstat-Dx
Analyzer 1.0. Do not use a QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge that is past its
expiration date, appears damaged, or leaks fluid. Dispose of used or damaged cartridges in
accordance with all national, state, and local health and safety regulations and laws.
Observe standard laboratory procedures for keeping the working area clean and
contamination-free. Guidelines are outlined in publications such as the
Microbiological and Biomedical Laboratories
from the Centers for Disease Control and
Biosafety in
Prevention and the National Institutes of Health (https://www.cdc.gov/labs/BMBL.html).

18
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Highly flammable liquid and vapour. Harmful if swallowed or if
ory tract. Keep away from
dust/fume/gas/mist/vapours/spray. Wear protective
gloves/protective clothing/eye protection/face protection. Wear
Continue rinsing. IF exposed or concerned: Immediately call a
and keep comfortable for breathing.
The following hazard and precautionary statements apply to components of the QIAstat-Dx
Respiratory SARS-CoV-2 Panel.
QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge
Contains: ethanol; guanidine hydrochloride; guanidine thiocyanate;
isopropanol; proteinase K; t-Octylphenoxypolyethoxyethanol. Danger!
inhaled. May be harmful in contact with skin. Causes severe skin burns
and eye damage. May cause allergy or asthma symptoms or breathing
difficulties if inhaled. May cause drowsiness or dizziness. Harmful to
aquatic life with long lasting effects. Contact with acids liberates very
toxic gas. Corrosive to the respirat
heat/sparks/open flames/hot surfaces. No smoking. Avoid breathing
respiratory protection. IF IN EYES: Rinse cautiously with water for
several minutes. Remove contact lenses, if present and easy to do.
POISON CENTER or doctor/ physician. Remove person to fresh air

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
19
Reagent Storage and Handling
Store the QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridges in a dry, clean storage space
at room temperature (15–25°C). Do not remove the QIAstat-Dx Respiratory SARS-CoV-2 Panel
Cartridges or the transfer pipettes from their individual packaging until actual use. Under these
conditions, QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridges can be stored until the
expiration date printed on the individual packaging. The expiration date is also included in
the QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge bar code and is read by the
QIAstat-Dx Analyzer 1.0 when the cartridge is inserted into the instrument to run a test.
Specimen Handling, Storage and Preparation
Nasopharyngeal samples should be collected and handled according to the manufacturer’s
recommended procedures.
Recommended storage conditions for NPS (nasopharyngeal swab) resuspended in UTM
specimens are listed below:
Room temperature up to 4 hours at 15–25°C
Refrigerated up to 3 days at 2–8°C
Frozen up to 30 days at –15 to –25°C

20
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Procedure
Internal Control
The QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge includes a full process Internal Control
which is titered MS2 bacteriophage. The MS2 bacteriophage is a single-stranded RNA virus
that is included in the cartridge in dried form and is rehydrated upon sample loading. This
Internal Control material verifies all steps of the analysis process, including sample
resuspension/homogenization, lysis, nucleic acid purification, reverse transcription and PCR.
A positive signal for the Internal Control indicates that all processing steps performed by the
QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge were successful.
A negative signal of the Internal Control does not negate any positive results for detected and
identified targets, but it does invalidate all negative results in the analysis. Therefore, the test
should be repeated if the Internal Control signal is negative.

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
21
Protocol: Transport medium liquid samples
Sample collection, transport and storage
Collect nasopharyngeal swab samples according to the swab manufacturer’s recommended
procedures and place the swab into Universal Transport Medium.
Loading a sample into the QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge
1. Open the package of a QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge using the
tear notches on the sides of the packaging (Figure 4).
IMPORTANT: After the package is open, sample should be introduced inside the
QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge and loaded into the QIAstat-Dx
Analyzer 1.0 within 120 minutes.
Figure 4. Opening the QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge.
2. Remove the QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge from the packaging and
position it so that the QR code on the label faces you.

22
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
3. Manually write the sample information or place a sample information label on the top of
the QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge. Make sure that the label is
properly positioned and does not block the lid opening (Figure 5).
Figure 5. Sample information placement on top of QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge.
4. Open the sample lid of the main port on the front of the QIAstat-Dx Respiratory
SARS-CoV-2 Panel Cartridge (Figure 6).
Figure 6. Opening the sample lid of main port.
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
23
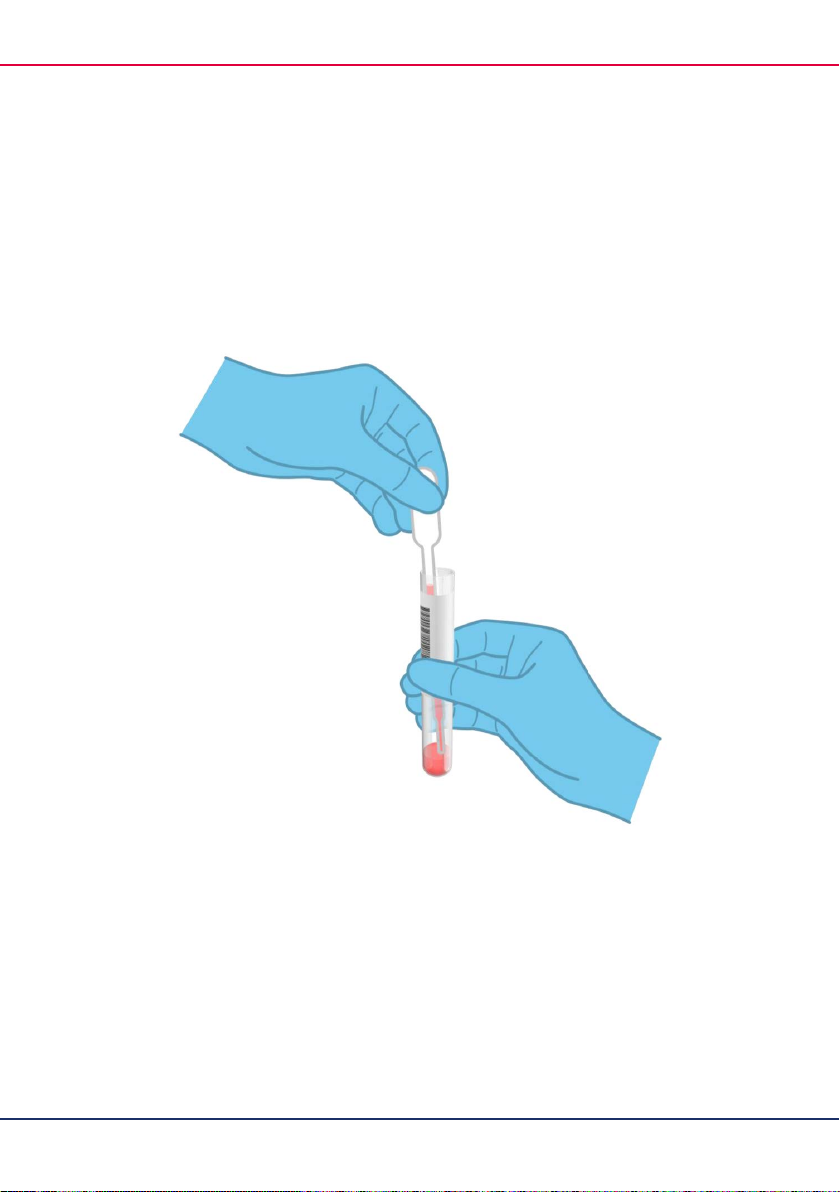
5. Open the tube with the sample to be tested. Use the supplied transfer pipette to draw up
fluid to the third fill line on the pipette (i.e., 300 μl) (Figure 7).
®
IMPORTANT: Take care to avoid drawing air into the pipette. If Copan
UTM®, Universal
Transport Medium is used as transport medium take care not to aspirate any of the beads
present in the tube. If air or beads are drawn into the pipette, carefully expel the sample
fluid in the pipette back into the sample tube and draw up fluid again.
Figure 7. Drawing up sample into the supplied transfer pipette.
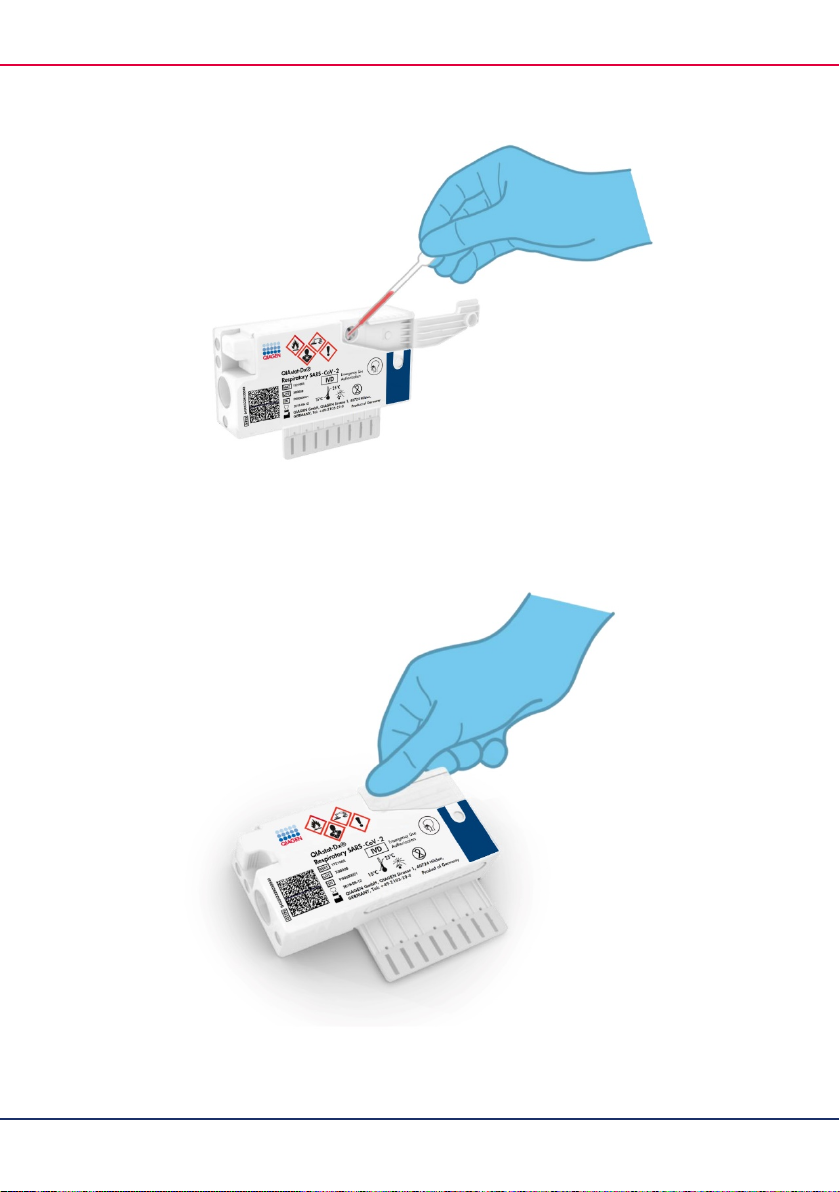
6. Carefully transfer 300 μl of sample volume into the main port of the QIAstat-Dx
Respiratory SARS-CoV-2 Panel Cartridge using the supplied single-use transfer pipette
(Figure 8, next page).
24
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Figure 8. Transferring sample to main port of QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge.
7. Firmly close the sample lid of the main port until it clicks (Figure 9).
Figure 9. Closing the sample lid of the main port.

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
25
8. Visually confirm that the sample has been loaded by checking the sample inspection
window of the QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge (Figure 10).
IMPORTANT: After the sample is placed inside the QIAstat-Dx Respiratory SARS-CoV-2
Panel Cartridge, the cartridge must be loaded into the QIAstat-Dx Analyzer 1.0 within
90 minutes.
Figure 10. Sample inspection window (blue arrow).
Starting the QIAstat-Dx Analyzer 1.0
9. Power ON the QIAstat-Dx Analyzer 1.0 using the On/Off button on the front of the
instrument.
Note: The power switch on the back of the Analytical Module must be set in the “I”
position. The QIAstat-Dx Analyzer 1.0 status indicators will turn blue.
10. Wait until the Main screen appears and the QIAstat-Dx Analyzer 1.0 status indicators
turn green and stop blinking.
11. Log in to the QIAstat-Dx Analyzer 1.0 by entering the user name and password.
Note: The Login screen will appear if User Access Control is activated. If the User Access Control
is disabled, no user name/password will be required and the Main screen will appear.

26
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
12. If the Assay Definition File software has not been installed on the QIAstat-Dx
Analyzer 1.0, follow the installation instructions prior to running the test (see “Appendix
A: Installing the Assay Definition File”, page 111, for additional information).
Running a test
13. Press the Run Test button in the top right corner of the touchscreen of the QIAstat-Dx
Analyzer 1.0.
14. When prompted, scan the sample ID bar code on the UTM tube containing the sample,
or scan the specimen information bar code located on the top of the QIAstat-Dx
Respiratory SARS-CoV-2 Panel Cartridge (see step 3) using the integrated front bar code
reader of the QIAstat-Dx Analyzer 1.0 (Figure 11).
Note: It is also possible to enter the sample ID using the virtual keyboard of the
touchscreen by selecting the Sample ID field.
Note: Depending on the chosen system configuration, entering the patient ID may also
be required at this point.
Note: Instructions from the QIAstat-Dx Analyzer 1.0 appear in the Instructions Bar at the
bottom of the touchscreen.
Figure 11. Scanning sample ID bar code.
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
27
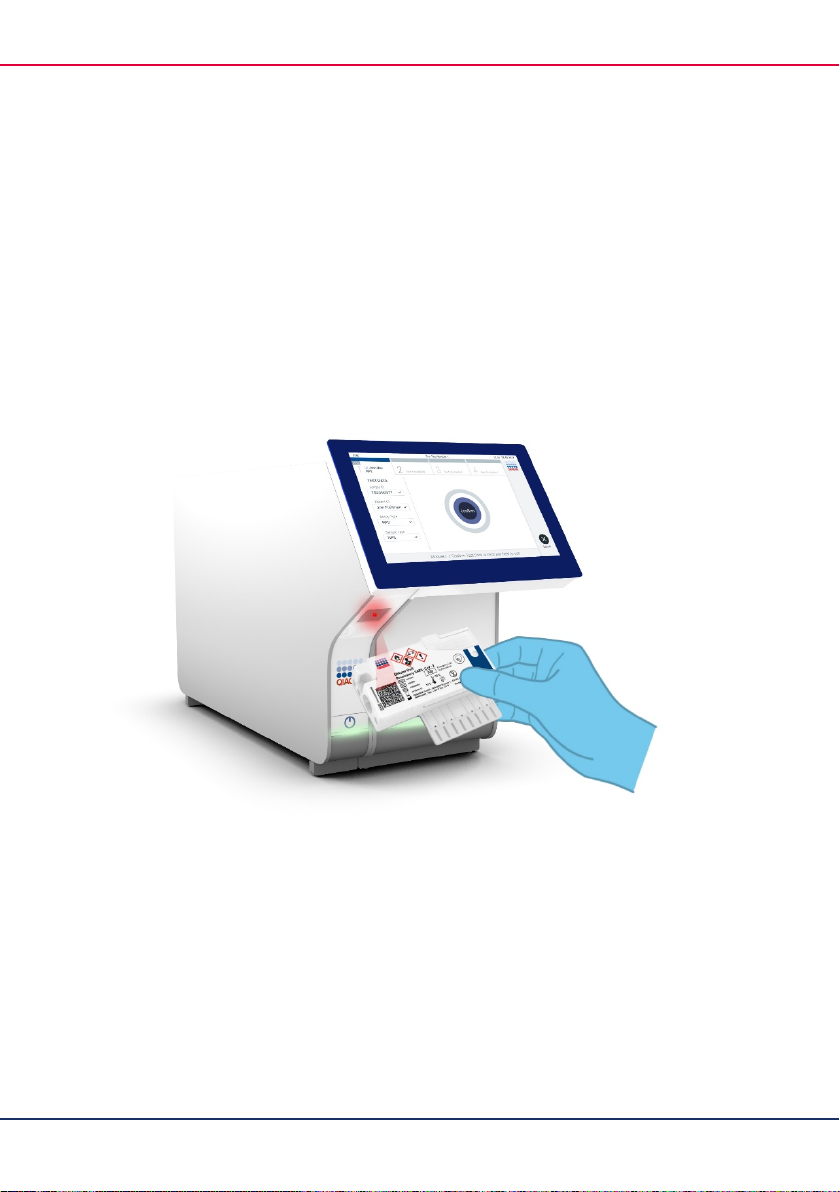
15. When prompted, scan the bar code of the QIAstat-Dx Respiratory SARS-CoV-2 Panel
Cartridge to be used (Figure 12). The QIAstat-Dx Analyzer 1.0 automatically recognizes
the assay to be run based on the cartridge bar code.
Note: The QIAstat-Dx Analyzer 1.0 will not accept QIAstat-Dx Respiratory SARS-CoV-2 Panel
Cartridges with lapsed expiration dates, previously used cartridges, or cartridges for assays
that have not been installed on the unit. An error message will be shown in these cases and
the QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge will be rejected. Refer to the
QIAstat-Dx Analyzer 1.0 User Manual
for further details on how to install assays.
Figure 12. Scanning QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge bar code.
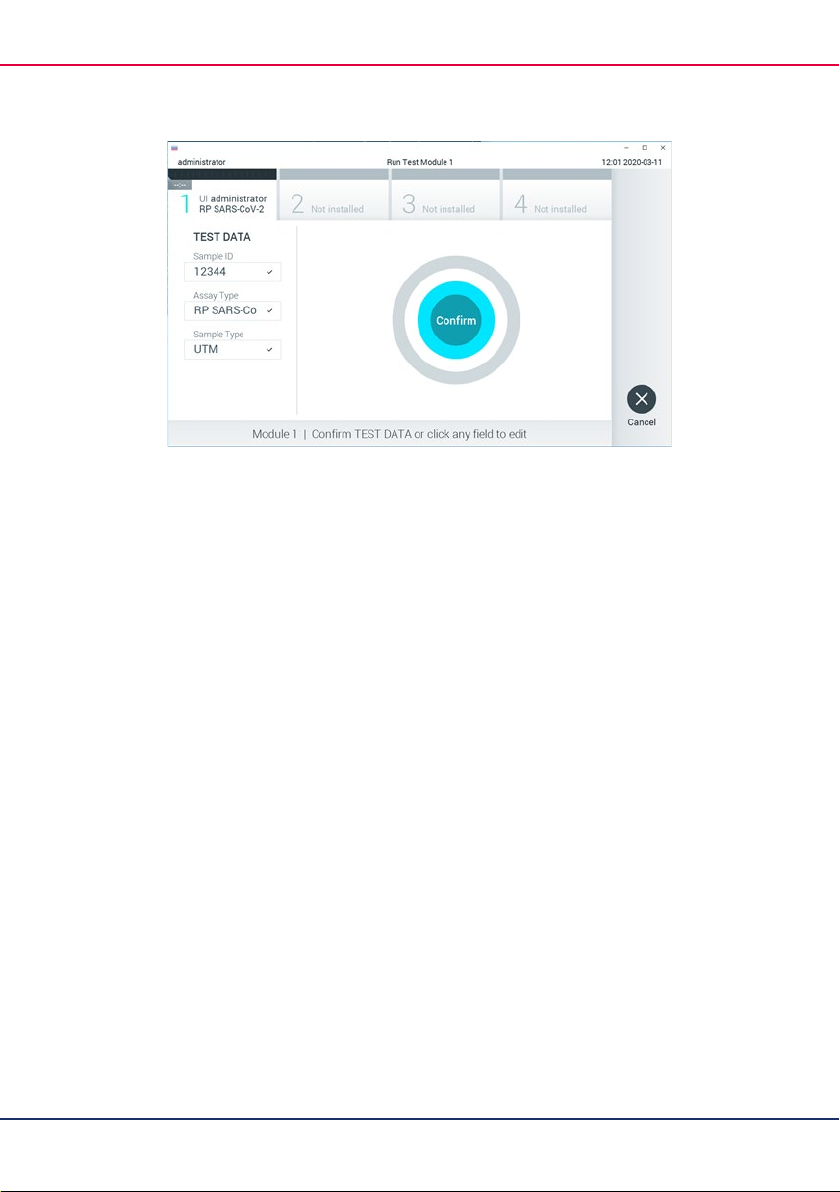
16. The Confirm screen will appear. Review the entered data and make any necessary
changes by selecting the relevant fields on the touchscreen and editing the information.
17. Press Confirm when all the displayed data are correct. If needed, select the appropriate
field to edit its content, or press Cancel to cancel the test (Figure 13, next page).
28
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Figure 13. Confirming data entry.
18. Make sure that both sample lids of the swab port and main port of the QIAstat-Dx
Respiratory SARS-CoV-2 Panel Cartridge are firmly closed. When the cartridge entrance
port on the top of the QIAstat-Dx Analyzer 1.0 automatically opens, insert the QIAstat-Dx
Respiratory SARS-CoV-2 Panel Cartridge with the bar code facing to the left and the
reaction chambers facing down (Figure 14, next page).
Note: There is no need to push the QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge
into the QIAstat-Dx Analyzer 1.0. Position it correctly into the cartridge entrance port and
the QIAstat-Dx Analyzer 1.0 will automatically move the cartridge into the Analytical
Module.

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
29
Figure 14. Inserting QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge into QIAstat-Dx Analyzer 1.0.

30
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
19. Upon detecting the QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge, the QIAstat-Dx
Analyzer 1.0 will automatically close the lid of the cartridge entrance port and start the
test run. No further action from the operator is required to start the run.
Note: The QIAstat-Dx Analyzer 1.0 will not accept a QIAstat-Dx Respiratory SARS-CoV-2
Panel Cartridge other than the one used and scanned during the test setup. If a cartridge
other than the one scanned is inserted, an error will be generated and the cartridge will
be automatically ejected.
Note: Up to this point, it is possible to cancel the test run by pressing the Cancel button in
the bottom right corner of the touchscreen.
Note: Depending on the system configuration, the operator may be required to re-enter
their user password to start the test run.
Note: The lid of the cartridge entrance port will close automatically after 30 seconds if a
QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge is not positioned in the port. If this
occurs, repeat the procedure starting with step 17.
20. While the test is running, the remaining run time is displayed on the touchscreen.
21. After the test run is completed, the Eject screen will appear (Figure 15, next page) and
the Module status bar will display the test result as one of the following options:
TEST COMPLETED: The test was completed successfully
TEST FAILED: An error occurred during the test
TEST CANCELED: The user canceled the test
IMPORTANT: If the test fails, refer to the “Troubleshooting” section in the
Analyzer 1.0 User Manual
for possible reasons and instructions on how to proceed.
QIAstat-Dx

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
31
Figure 15. Eject screen display.
22. Press Eject on the touchscreen to remove the QIAstat-Dx Respiratory SARS-CoV-2
Panel Cartridge and dispose of it as biohazardous waste in accordance with all
national, state, and local health and safety regulations and laws. The QIAstat-Dx
Respiratory SARS-CoV-2 Panel Cartridge should be removed when the cartridge entrance
port opens and ejects the cartridge. If the cartridge is not removed after 30 seconds, it
will automatically move back into the QIAstat-Dx Analyzer 1.0 and cartridge entrance
port lid will close. If this occurs, press Eject to open the lid of the cartridge entrance port
again and then remove the cartridge.
IMPORTANT: Used QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridges must be
discarded. It is not possible to re-use cartridges for tests for which the execution was
started but then subsequently cancelled by the operator, or for which an error was
detected.
23. After the QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge has been ejected, the
results Summary screen will appear. Refer to “Interpretation of Results”, page, 34 for
further details. To begin the process for running another test, press Run Test.
Note: For further information on the use of the QIAstat-Dx Analyzer 1.0, refer to the
QIAstat-Dx Analyzer 1.0 User Manual
.

32
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Viewing results
The QIAstat-Dx Analyzer 1.0 automatically interprets and saves test results. After ejecting the
QIAstat-Dx Respiratory SARS-CoV-2 Panel Cartridge, the results Summary screen is
automatically displayed (Figure 16).
Figure 16. Results Summary screen example showing Test Data on the left panel and Test Summary in the main panel.
The main part of the screen provides the following three lists and uses color-coding and
symbols to indicate the results:
The first list includes all pathogens detected and identified in the sample, preceded by a
sign and are colored red.
The second list includes all equivocal pathogens, preceded by a yellow question
mark , in the event any of the subtypes H1, H3 and/or H1N1 pdm09 are detected
and identified in the sample, but Influenza A is not detected.
The third list includes all pathogens tested in the sample. Pathogens detected and
identified in the sample are preceded by a sign and are colored red. Pathogens that
were tested but not detected are preceded by a sign and are colored green.
Equivocal pathogens are preceded by a .
Note: Pathogens detected and identified in the sample are shown in all lists.

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
33
If the test failed to complete successfully, a message will indicate “Failed”, followed by the
specific Error Code.
The following Test Data is shown on the left side of the screen:
Sample ID
Assay Type
Sample Type
Further data about the assay is available, depending on the operator’s access rights, through
the tabs at the bottom of the screen (e.g., amplification plots and test details). For additional
details, please see section below.

34
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Interpretation of Results
Internal Control interpretation
Internal Control results are to be interpreted according to Table 2.
Table 2. Interpretation of Internal Control results
Control result Explanation Action
Passed The Internal Control amplified
successfully
Failed The Internal Control failed
Pathogen Result interpretation
The run was completed with success. All results
are validated and can be reported. Detected
pathogens are reported as “positive” and
undetected pathogens are reported as “negative”.
Positively detected pathogen(s) are reported, but
all negative results (tested but not detected
pathogen[s]) are invalid.
Repeat the testing using a new QIAstat-Dx
Respiratory SARS-CoV-2 Panel Cartridge.
Result interpretation information for SARS-CoV-2:
The SARS-CoV-2 in this panel targets two genes of the virus genome (Orf1b poly gen
(Rdrp gene) and E genes) detected with the same fluorescence channel. The two targets are
not differentiated, and amplification of either or both regions leads to a fluorescence signal.
Result Interpretation information for Influenza A
A result for a respiratory organism is interpreted as “Positive” when the corresponding PCR
assay is positive (see exceptions for Influenza A below). The Influenza A assay in the QIAstatDx Respiratory SARS-CoV-2 Panel is designed to detect Influenza A as well as Influenza A
subtype H1N1/2009, Influenza A subtype H1, or Influenza A subtype H3. In particular, this
means:

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
35
If seasonal Influenza A H1 strain is detected by the QIAstat-Dx Respiratory SARS-CoV-2
Panel assay, two signals will be generated and displayed on the QIAstat-Dx Analyzer
1.0 screen: one for Influenza A and a second one for H1 strain.
Note: It is acceptable if only the H1 signal is obtained, which would be indicated as
“equivocal”.
If seasonal Influenza A H3 strain is detected by the QIAstat-Dx Respiratory SARS-CoV-2
Panel assay, two signals will be generated and displayed on the QIAstat-Dx Analyzer
1.0 screen: one for Influenza A and a second one for H3 strain.
Note: It is acceptable if only the H3 signal is obtained, which would be indicated as
“equivocal”.
If a pandemic Influenza A/H1N1/2009 strain is detected, two signals will be generated
and displayed on the QIAstat-Dx Analyzer 1.0 screen: one for Influenza A and a second
one for H1N1/2009.
Note: It is acceptable if only the H1N1/2009 signal is obtained, which would be
indicated as “equivocal”.
Note: It is acceptable if only the Influenza A signal is obtained, which would be
indicated as “Influenza A (no subtype detected)”.
IMPORTANT: If only an Influenza A signal is present and no additional signal for any of the
subtypes is generated, it can be due to either low concentration or, in very rare cases, a new
variant or any Influenza A strain other than H1 and H3 (eg, H5N1, which can infect humans).
See important precautions regarding possible detection of Influenza A with no subtype
detected.
Result Interpretation for all other pathogens
For every other pathogen that can be detected with the QIAstat-Dx Respiratory SARS-CoV-2
Panel, only one signal will be generated if the pathogen is present in the sample.

36
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Viewing amplification curves
To view test amplification curves of pathogens detected, press the Amplification Curves tab
(Figure 17).
Figure 17. Amplification Curves screen (PATHOGENS tab).
Details about the tested pathogens and controls are shown on the left, and the amplification
curves are shown in the center.
Note: If User Access Control is enabled on the QIAstat-Dx Analyzer 1.0, the Amplification
Curves screen is only available for operators with access rights.
Press the PATHOGENS tab on the left side to display the plots corresponding to the tested
pathogens. Press on the pathogen name to select which pathogens are shown in the
amplification plot. It is possible to select single, multiple, or no pathogens. Each pathogen in
the selected list will be assigned a color corresponding to the amplification curve associated
with the pathogen. Unselected pathogens will be shown in gray.
The corresponding C
and endpoint fluorescence (EP) values are shown below each pathogen
T
name.

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
37
Press the CONTROLS tab on the left side to view the controls in the amplification plot. Press the
circle next to the control name to select or deselect it (Figure 18).
Figure 18. Amplification Curves screen (CONTROLS tab).
The amplification plot displays the data curve for the selected pathogens or controls. To
alternate between logarithmic or linear scale for the Y-axis, press the Lin or Log button at the
bottom left corner of the plot.
The scale of the X-axis and Y-axis can be adjusted using the blue pickers on each axis.
Press and hold a blue picker and then move it to the desired location on the axis. Move a blue
picker to the axis origin to return to the default values.
Viewing test details
Press Test Details in the Tab Menu bar at the bottom of the touchscreen to review the results
in more detail. Scroll down to see the complete report. The following Test Details are shown
in the center of the screen (Figure 19, next page):
User ID
Cartridge SN (serial number)

38
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Cartridge Expiration Date
Module SN (serial number)
Test Status (Completed, Failed, or Canceled by operator)
Error Code (if applicable)
Test Start Date and Time
Test Execution Time
Assay Name
Test ID
Test Result:
Positive (if at least one respiratory pathogen is detected/identified)
Positive with warning (at least one respiratory pathogen is detected but the Internal
Control failed)
Negative (no respiratory pathogen is detected)
Invalid
List of analytes tested in the assay, with C
and endpoint fluorescence in the event of a
T
positive signal
Internal Control, with C
Figure 19. Example screen showing Test Data on the left panel and Test Details in the main panel.
and endpoint fluorescence
T

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
39
Browsing results from previous tests
To view results from previous tests that are stored in the results repository, press View Results
on the Main Menu bar (Figure 20).
Figure 20. Example View Results screen.
The following information is available for every executed test (Figure 21):
Sample ID
Assay (name of test assay)
Operator ID
Mod (Analytical Module on which the test was executed)
Date/Time (date and time when the test was finished)
Result (outcome of the test: positive [pos], positive with warning [pos*], negative [neg],
failed [fail] or successful [suc])
Note: If User Access Control is enabled on the QIAstat-Dx Analyzer 1.0, the data for which
the user has no access rights will be hidden with asterisks.

40
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Select one or more test results by pressing the gray circle to left of the sample ID. A checkmark
will appear next to selected results. Unselect test results by pressing this checkmark. The entire
list of results can be selected by pressing the checkmark circle in the top row (Figure 21).
Figure 21. Example of selecting Test Results in the View Results screen.
Press anywhere in the test row to view the result for a particular test.
Press a column headline (e.g., Sample ID) to sort the list in ascending or descending order
according to that parameter. The list can be sorted according to only one column at a time.
The Result column shows the outcome of each test (Table 3):

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
41
Table 3. Descriptions of test results
Outcome Result Description
Positive
Positive with warning
Negative
Failed
Successful
pos
pos*
neg
fail
suc
At least one pathogen is positive.
At least one pathogen is positive but the Internal Control failed.
No pathogens were detected.
The test failed because either an error occurred or the test was
canceled by the user.
The test is either positive or negative, but the user does not have
the access rights to view the test results
Make sure a printer is connected to the QIAstat-Dx Analyzer 1.0 and the proper driver is
installed. Press Print Report to print the report(s) for the selected result(s).
Press Save Report to save the report(s) for the selected result(s) in PDF format to an external
USB storage device.
Select the report type: List of Tests or Test Reports.
Press Search to search the test results by Sample ID, Assay and Operator ID. Enter the search
string using the virtual keyboard and press Enter to start the search. Only the records
containing the search text will be displayed in the search results.
If the results list has been filtered, the search will only apply to the filtered list.
Press and hold a column headline to apply a filter based on that parameter. For some
parameters, such as Sample ID, the virtual keyboard will appear so the search string for the
filter can be entered.
For other parameters, such as Assay, a dialog will open with a list of assays stored in the
repository. Select one or more assays to filter only the tests that were performed with the
selected assays.

42
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
The symbol to the left of a column headline indicates that the column’s filter is active.
A filter can be removed by pressing Remove Filter in the Submenu bar.
Exporting results to a USB drive
From any tab of the View Results screen, select Save Report to export and save a copy of the
test results in PDF format to a USB drive. The USB port is located on the front of the QIAstat-Dx
Analyzer 1.0.
Printing results
Make sure a printer is connected to the QIAstat-Dx Analyzer 1.0 and the proper driver is
installed. Press Print Report to send a copy of the test results to the printer.

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
43
Quality Control
In accordance with QIAGEN’s ISO-certified Quality Management System, each lot of
QIAstat-Dx Respiratory SARS-CoV-2 Panel is tested against predetermined specifications to
ensure consistent product quality. External controls are not provided with the QIAstat-Dx
Respiratory SARS-CoV-2 Panel. Quality control requirements should be performed in
conformance with local, state, and/or federal regulations or accreditation requirements and
your laboratory’s standard quality control procedures.
The following external controls are available:
ZeptoMetrix
ZeptoMetrix Inc. NATtrol SARS-CoV-2 E/ORF1ab recombinant (Cat. No. 0831043)
®
Inc. NATtrol™ Respiratory Verification Panel (Cat. No. NATRVP-QIA)

44
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Limitations
For prescription use only.
The use of this assay as an in vitro diagnostic under FDA Emergency Use Authorization
(EUA) is limited to laboratories that are certified under the Clinical Laboratory
Improvement Amendments of 1988 (CLIA), 42 U.S.C. §263a, to perform high and
moderate complexity tests.
Results from the QIAstat-Dx Respiratory SARS-CoV-2 Panel are not intended to be used as
the sole basis for diagnosis, treatment, or other patient management decisions.
Laboratories are required to report all positive SARS-CoV-2 results to the appropriate
public health authorities.
Primers and probes for this kit target highly conserved regions within the genome of
SARS- CoV-2. Mutations occurring in these highly conserved regions (although rare) may
result in RNA being undetectable.
The E gene target is homologous to sequences from multiple bat SARS viruses. These
viruses are unlikely to be found in nasopharyngeal swabs and their potential to infect
human hosts is unknown.
The performance of this test has not been established for immunocompromised
individuals.
The performance of this test has not been established for patients without signs and
symptoms of respiratory infection.
Positive results do not rule out co-infection with organisms not included in the QIAstat-Dx
Respiratory SARS-CoV-2 Panel. The agent detected may not be the definitive cause of the
disease.
Negative results do not preclude infection of the upper respiratory tract. Not all agents of
acute respiratory infection are detected by this assay and sensitivity in some clinical
settings may differ from that described in the package insert.

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
45
A negative result with the QIAstat-Dx Respiratory SARS-CoV-2 Panel does not exclude the
infectious nature of the syndrome. Negative assay results may originate from several
factors and their combinations, including sample handling mistakes, variation in the
nucleic acid sequences targeted by the assay, infection by organisms not included in the
assay, organism levels of included organisms that are below the limit of detection for the
assay, and use of certain medications, therapies, or agents.
The QIAstat-Dx Respiratory SARS-CoV-2 Panel is not intended for testing of samples other
than those described in these Instructions for Use. Test performance characteristics have
been established only with nasopharyngeal swab samples collected in universal transport
media (UTM) from individuals with acute respiratory symptoms.
The QIAstat-Dx Respiratory SARS-CoV-2 Panel is intended to be used in conjunction with
standard of care culture for organism recovery, serotyping, and/or antimicrobial
susceptibility testing where applicable.
The results from the QIAstat-Dx Respiratory SARS-CoV-2 Panel must be interpreted by a
trained healthcare professional within the context of all relevant clinical, laboratory, and
epidemiological findings.
The QIAstat-Dx Respiratory SARS-CoV-2 Panel can be used only with the QIAstat-Dx
Analyzer 1.0.
The QIAstat-Dx Respiratory SARS-CoV-2 Panel is a qualitative assay and does not
provide a quantitative value for detected organisms.
Viral and bacterial nucleic acids may persist in vivo, even if the organism is not viable or
infectious. Detection of a target marker does not imply that the corresponding organism
is the causative agent of the infection or the clinical symptoms.
Detection of viral and bacterial nucleic acids depends on proper sample collection,
handling, transportation, storage and loading into the QIAstat-Dx Respiratory
SARS-CoV-2 Panel Cartridge. Improper operations for any of the aforementioned
processes can cause incorrect results, including false-positive or false-negative results.
The performance of this test has not been established for screening of blood or blood products.

46
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
The performance of this test has not been established in individuals who received
influenza vaccine. Recent administration of a nasal influenza vaccine may cause false
positive results for Influenza A and/or Influenza B.
The QIAstat-Dx Respiratory SARS-CoV-2 Panel may not be able to distinguish between
existing viral strains and new variants as they emerge. For example, the QIAstat-Dx
Respiratory SARS-CoV-2 Panel can detect seasonal H3N2 Influenza but may not be able
to distinguish seasonal H3N2 from H3N2 variant (H3N2v).
The QIAstat-Dx Respiratory SARS-CoV-2 Panel detects the multi-copy IS481 insertion
sequence present in multiple
possible if the specimen is contaminated with non-pertussis
The assay sensitivity and specificity, for the specific organisms and for all organisms
combined, are intrinsic performance parameters of a given assay and do not vary
depending on prevalence. In contrast, both the negative and positive predictive values of
a test result are dependent on the disease/organism prevalence. False negative test
results are more likely during peak activity when prevalence of disease is high. False
positive test results are more likely during periods when prevalence is moderate or low.
Bordetella
species. False positive
Bordetella
B. pertussis
species.
results are

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
47
Conditions of Authorization for the Laboratory
The QIAstat-Dx Respiratory SARS-CoV-2 Panel Letter of Authorization, along with the
authorized Fact Sheet for Healthcare Providers, the authorized Fact Sheet for Patients, and
authorized labeling are available on the FDA website:
https://www.fda.gov/MedicalDevices/Safety/EmergencySituations/ucm161496.htm.
However, to assist clinical laboratories using the QIAstat-Dx Respiratory SARS-CoV-2 Panel
(“your product” in the conditions below), the relevant Conditions of Authorization are listed
below:
Authorized laboratories * using your product will include with result reports of your
product, all authorized Fact Sheets. Under exigent circumstances, other appropriate
methods for disseminating these Fact Sheets may be used, which may include mass
media.
Authorized laboratories using your product will use your product as outlined in the
Instructions for Use. Deviations from the authorized procedures, including the authorized
instruments, authorized extraction methods, authorized clinical specimen types,
authorized control materials, authorized other ancillary reagents, and authorized
materials required to use your product are not permitted.
Authorized laboratories that receive your product will notify the relevant public health
authorities of their intent to run your product prior to initiating testing.
Authorized laboratories using your product will have a process in place for reporting test
results to healthcare providers and relevant public health authorities, as appropriate.
* The letter of authorization refers to, “United States (U. S.) laboratories certified under the Clinical Laboratory
Improvement Amendments of 1988 (CLIA), 42 U.S.C. §263a, to perform high and moderate complexity tests” as
“authorized laboratories.”

48
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Authorized laboratories will collect information on the performance of your product and
will report to DMD/OHT7-OIR/OPEQ/CDRH (via email: CDRH-EUAReporting@fda.hhs.gov) and QIAGEN (https://www.qiagen.com/us/service-andsupport/technical-support/technical-support-form/) any suspected occurrence of false
positive or false negative results, and significant deviations from the established
performance characteristics of your product of which they become aware.
All laboratory personnel using your product must be appropriately trained in RT-PCR
techniques, use appropriate laboratory and personal protective equipment when
handling this kit, and use your product in accordance with the authorized labeling.

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
49
Performance Characteristics
The QIAstat-Dx Respiratory SARS-CoV-2 Panel (Cat. No. 691223) assay was developed by
introducing the reagents required to detect the SARS-CoV-2 target in a separate reaction
chamber of the QIAstat-Dx Respiratory Panel assay (Cat. No. 691221), leaving all other
targets unchanged. As a result of this and/or availability of SARS-CoV-2 clinical samples,
certain studies shown below were not done or repeated using the QIAstat-Dx Respiratory
SARS-CoV-2 Panel.
SARS-CoV-2 Target
Clinical performance
The performance of SARS-CoV-2 target in the QIAstat-Dx Respiratory SARS-CoV-2 Panel was
evaluated using retrospective nasopharyngeal swab clinical specimens in transport medium.
Specifically, 30 individual negative nasopharyngeal specimens and a total of 30 positive
samples consisting of:
10 positive clinical samples tested with a validated molecular comparator assay
obtained from a Hospital in Barcelona (Spain)
20 contrived positive clinical samples at 1-2x LoD
All clinical samples were collected from patients with signs and symptoms of upper respiratory
infection by qualified personnel according to the package insert of the collection device and
stored frozen until use.
Low positive contrived clinical samples were prepared by spiking a quantified clinical sample
obtained from a Hospital in Barcelona (Spain) into individual negative clinical samples to
approximately 1x- 2x LoD (20 samples).

50
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Overall results are shown on Table 4:
Table 4. Overall clinical performance results of SARS-CoV-2 target
Sample Sample Type N
Positives Positive clinical sample 10 (10/10) 100% N/A 0/0 N/A
Total Positive Samples 30 (30/30) 100% 85.8 -100 0/0 N/A
Negative Total Negative Samples 30 0/0 N/A (30/30) 100% 85.8 -100
Low positive contrived
sample (1x-2xLoD)
% Positive (95% CI) % Negative
(20/20) 100% N/A 0/0 N/A
20
SARS-CoV-2 Target
(95% CI)
Performance of the SARS-CoV-2 target in the QIAstat-Dx Respiratory SARS-CoV-2 Panel against
the expected results are:
Positive Percent Agreement (PPA%): 30/30 = 100 % (95% CI: 85.8% - 100%)
Negative Percent Agreement (NPA%): 30/30 = 100% (95% CI: 85.8% - 100%)
Analytical performance
Sensitivity (Limit of Detection)
The Analytical Sensitivity, or Limit of Detection (LoD), is defined as the lowest concentration at
which ≥95% of the tested samples generate a positive call.
The LoD of the SARS-CoV-2 target was assessed by analyzing serial dilutions of analytical
samples prepared from a quantified clinical sample obtained from a Hospital from Barcelona
(Spain). Dilutions were preformed using simulated matrix consisting of UTM and HeLa cells.
Four (4) replicates were tested of each serial dilution. The lowest concentration at which all
replicates were positive was interpreted as the tentative LoD. The LoD was then confirmed by
testing twenty (20) replicates with concentrations at the tentative limit of detection. To confirm
the established LoD concentration, the detection rate of all replicates must be ≥95% (at least
19/20 replicates must generate a positive signal).

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
51
Table 5. LoD value obtained for the SARS-CoV-2 target tested with the QIAstat-Dx Respiratory SARS-CoV-2 Panel
Pathogen Strain Source Concentration Detection rate
SARS-CoV-2 Clinical Sample Hospital from Barcelona (Spain) 500 copies/mL 20/20
The final LoD for the SARS-CoV-2 target in the QIAstat-Dx Respiratory SARS-CoV-2 panel
according to the assay results interpretation, is 500 copies/ml.
FDA SARS-CoV-2 reference panel testing
The evaluation of sensitivity and MERS-CoV cross-reactivity was performed using reference
material (T1), blinded samples, and a standard protocol provided by the FDA. The study
included a range-finding study and a confirmatory study for LoD. Blinded sample testing was
used to establish specificity and to confirm the LoD. All the reagents required for the complete
execution of the test are pre-loaded and self-contained in the QIAstat-Dx Respiratory
SARS-CoV-2 Panel cartridge. The instrument used was the QIAstat-Dx Analyzer 1.0. The results
are summarized in Table 6 .
Table 6. Summary of LoD confirmation result using the FDA SARS-CoV-2 Reference Panel
Reference Materials Provided by FDA Specimen Type
SARS-CoV-2
MERS-CoV N/A ND
Abbreviations: NDU/ml = RNA NAAT detectable units/ml; ND = not detected; N/A = not applicable.
Nasopharyngeal swab
Product LoD Cross-Reactivity
1.8x105 NDU/ml N/A
Exclusivity (Cross-reactivity and Exclusivity)
The analytical specificity study was carried out by
the cross-reactivity and exclusivity of the SARS-CoV-2 target. On-panel organisms were tested to
assess the potential for intra-panel cross-reactivity and off-panel organisms were tested to evaluate
panel exclusivity. The off-panel organisms selected were clinically relevant organisms (colonizing
the upper respiratory tract or causing respiratory symptoms), common skin flora, or laboratory
contaminants, or microorganisms for which much of the population may have been infected.
* Only a limited number of organisms were tested
in vitro
in silico
analysis and
(shown in Table 7).
in vitro *
testing to assess

52
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Samples were prepared by spiking potential cross-reactive organisms into simulated
nasopharyngeal swab sample matrix at the highest concentration possible based on the
5
organism stock – at least 10
TCID50/ml for viral targets and 106 CFU/ml for bacterial and
fungal targets. These concentrations represent levels approximately 800–1,000,000-fold
higher than the LoD of the SARS-CoV-2 Target.
Table 7. List of Analytical Specificity pathogens tested
Type Pathogen
On-panel bacteria
Off-panel bacteria
Off-panel viruses
* SARS Coronavirus was tested using custom gBlocks from the two regions targeted by the SARS-CoV-2 designs.
In silico
, sequence hits were analyzed together in order to detect unique specific sequences
in vitro
Chlamydophila pneumoniae
Haemophilus influenzae
Streptococcus pyogenes
Streptococcus pneumoniae
Mycobacterium tuberculosis
MERS Coronavirus
SARS Coronavirus*
matching with all primers and probes to be considered as positive amplifications. Primers and
probes were considered as reactive if the following parameters were fulfilled:
At least one forward, one probe, and one reverse primer of the SARS-CoV-2 assay match
with the target BLAST hit sequence.
At least 70% of query cover/identity between the BLAST hit sequence and every single
primer/probe sequence.
A maximum of 500 bp of amplicon size.
This analysis of the SARS-CoV-2 designs show that a potential unspecific signal can be
produced by a cross-reaction with a group of coronaviruses found in bats. These coronaviruses
have only been detected in bats and have not been reported to infect or colonize humans. No
unspecific signals were generated with critical off-panel human targets.

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
53
No cross-reactivity was observed, both
pathogens (colonizing the upper respiratory tract or causing respiratory symptoms), or
common sin flora or laboratory contaminants, or microorganisms.
in silico
and
in vitro
, with any clinically relevant
Inclusivity (Analytical Reactivity)
In silico
analysis shows that the SARS-CoV-2 assays in the QIAstat-Dx Respiratory SARS-CoV2
panel show a 100% sequence identity to 1184 out of the 1196 (99.0%) SARS-CoV-2 genomes
available in the public databases.
Twelve genome sequences showed a single nucleotide mismatch with one oligonucleotide
sequence of the E gene assay. These single mismatches are tolerated by the PCR workflow in
the QIAstat-Dx system and these genetic variants of the SARS-CoV-2 virus are predicted to be
detected by the QIAstat-Dx Respiratory SARS-CoV-2 panel despite the mismatches.
Additional Targets included in the QIAstat-Dx SARS-CoV-2 Panel
The performance of the other targets in the panel (Adenovirus, Coronavirus 229E, Coronavirus
HKU1, Coronavirus NL63, Coronavirus OC43, Human Metapneumovirus A+B, Influenza A,
Influenza A H1, Influenza A H3, Influenza A H1N1/pdm09, Influenza B, Parainfluenza virus 1,
Parainfluenza virus 2, Parainfluenza virus 3, Parainfluenza virus 4, Rhinovirus/Enterovirus,
Respiratory Syncytial Virus A+B,
Mycoplasma pneumoniae
) has been previously established and is presented below.
Bordetella pertussis, Chlamydophila pneumoniae
, and
Clinical performance
The clinical performance of the QIAstat-Dx Respiratory Panel (Cat. No. 691221) was
established during a multi-center study conducted at six (6) geographically diverse study sites:
five (5) U.S. sites and one (1) international site. Each study location was representative of the
intended use setting (clinical laboratories), and testing was performed by trained clinical
laboratory personnel. Residual nasopharyngeal swab (NPS) samples were collected from
subjects with signs and symptoms of respiratory infection for QIAstat-Dx Respiratory Panel and
comparator testing.

54
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Residual NPS specimens in UTM were tested with the QIAstat-Dx Respiratory Panel (Cat.
No. 691221) and an FDA-cleared molecular comparator, in accordance with product
instructions for use. Specimens tested in the clinical study were collected using the Universal
™
Transport Medium (UTM) (Copan Diagnostics [Brescia, Italy and CA, USA]), MicroTest
®
, M5®, M6® (Thermo Fisher Scientific®, MA, USA), BD™ Universal Viral Transport (UVT)
M4RT
System (Becton Dickinson, NJ, USA), Universal Transport Medium (UTM) System (HealthLink
®
Inc., FL, USA), Universal Transport Medium (Diagnostic Hybrids
®
(Quest Diagnostics
, NJ, USA) and UniTranz-RT® Universal Transport Media (Puritan®
, OH, USA), V-C-M Medium
M4®,
®
Diagnostics, ME, USA) collection kits.
A total of 2304 residual NPS specimens (1994 prospective, 310 archived) were tested in this
comparison study. Between December 2017 to April 2019, specimens were prospectively
collected from all comers meeting the study inclusion criteria and immediately frozen for later
testing by the study site as frozen prospective specimens (N=1093). No frozen samples were
distributed amongst sites. At time of testing, specimens were thawed and tested on both the
QIAstat-Dx Respiratory Panel and comparator method.
Between February and August 2018, specimens were prospectively collected from all comers
meeting the study eligibility criteria and tested fresh (N=901) on both the QIAstat-Dx Respiratory
Panel and comparator method in accordance with product instructions as fresh prospective
specimens. One specimen was withdrawn from the study due to an incorrect specimen type.
Table 8 (next page) provides the summary of demographic information for the 1994 subjects
that participated in the prospective study.

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
55
Table 8. Demographic summary for the prospective study arm
300
Site 1
Copenhagen,
Denmark
186 0 196 177 170 195
232 0 230 271 133 204
126 0 103 49 216 133
110 0 56 107 7 50
148 0 227 254 1 168
272 0 50 44 145 277
145 0 318 0 101 122
Overall
Male 924
(46.3%)
SEX
Female 1070
(53.7%)
≤5 years 627
(31.4%)
6–21 years 239
(11.9%)
AGE
22–49 years 330
(16.5%)
50+ years 798
(40.0%)
Outpatient 788
(39.5%)
Hospitalized 686
(34.4%)
Emergency 67
(3.4%)
ICU 153
(7.7%)
STATUS
Not
provided/
unknown
Total
(15.0%)
1994 418 0 426 448 303 399
Site 2
Minneapolis,
MN
Site 3
Indianapolis,
IN
Site 4
Liverpool,
NY
Site 5
Columbus,
OH
Site 6
Albuquerque,
NM
34 0 40 38 79 48
0 0 9 34 24 0
1 0 49 70 33 0
0 0 0 300 0 0

56
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
A total of 1994 specimens were evaluated for all panel members in the prospective study. The
performance of the QIAstat-Dx Respiratory Panel was evaluated by comparing the QIAstat-Dx
Respiratory Panel test results with those from an FDA-cleared multiplexed respiratory pathogen
panel.
Positive Percent Agreement (PPA) for each analyte was calculated as 100% x (TP/[TP+FN]).
True Positive (TP) indicates that both the QIAstat-Dx Respiratory Panel and the comparator
method yielded a “Detected” result of that specific analyte. A False Negative (FN) indicates
that the QIAstat-Dx Respiratory Panel was “Not Detected” while the comparator method was
“Detected” for the analyte in question. Negative Percent Agreement (NPA) was calculated as
100% x (TN/[TN+FP]). True Negative (TN) indicates that both the QIAstat-Dx Respiratory
Panel and the comparator method resulted in “Not Detected” for that specific analyte. A False
Positive (FP) indicates that the QIAstat-Dx Respiratory Panel was “Detected” while the
comparator method was “Not Detected” for the specific pathogen.
Binomial two-sided 95% Confidence Intervals were calculated using the Wilson Score Method.
The QIAstat-Dx Respiratory Panel prospective performance data in positive percent and
negative percent agreements against the comparator methods are presented by analyte in
Table 9, next page.

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
57
Table 9. QIAstat-Dx Respiratory Panel prospective clinical performance summary
Analyte TP/(TP+FN) Sensitivity/PPA 95% CI TN/(TN+FP) Specificity/NPA 95% CI
Viruses
Adenovirusa Fresh 55/58 94.8%
Coronavirus
229E
Coronavirus
HKU1b
Coronavirus
NL63
Coronavirus
OC43d
Frozen 31/32 96.9%
Overall 86/90 95.6%
Fresh 8/9 88.9%
Frozen 0/0 N/A N/A 1089/1089 100.0%
Overall 8/9 88.9% 56.5–
Fresh 3/3 100.0% 43.8–
Frozen 48/49 98.0% 89.3–
Overall 51/52 98.1% 89.9–
Fresh 4/5 80.0% 37.6–
c
Frozen 36/42 85.7% 72.2–
Overall 40/47 85.1% 72.3–
Fresh 3/3 100.0% 43.8–
Frozen 23/26 88.5% 71.0–
Overall 26/29 89.7% 73.6–
85.9–
98.2
84.3–
99.4
89.1–
98.3
56.5–
98.0
98.0
100.0
99.6
99.7
96.4
93.3
92.6
100.0
96.0
96.4
833/839 99.3%
1047/1057 99.1%
1880/1896 99.2%
886/886 100.0%
1975/1975 100.0% 99.8–
890/892 99.8% 99.2–
1035/1040 99.5% 98.9–
1925/1932 99.6% 99.3–
890/890 100.0% 99.6–
1046/1048 99.8% 99.3–
1936/1938 99.9% 99.6–
892/892 100.0% 99.6–
1059/1063 99.6% 99.0–
1951/1955 99.8% 99.5–
(continued on next page)
98.4–
99.7
98.3–
99.5
98.6–
99.5
99.6–
100.0
99.6–
100.0
100.0
99.9
99.8
99.8
100.0
99.9
100.0
100.0
99.9
99.9

58
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Table 9 (continued)
Analyte TP/(TP+FN) Sensitivity/PPA 95% CI TN/(TN+FP) Specificity/NPA 95% CI
Viruses
(continued)
Human
Fresh 62/67 92.5%
Metapneumoviruse
Frozen 53/55 96.4%
Overall 115/122 94.3%
Rhinovirus/
Enterovirus
Fresh 144/157 91.7%
f
Frozen 124/137 90.5%
Overall 268/294 91.2%
Influenza Ag Fresh 132/133 99.2%
Frozen 110/111 99.1% 95.1–
Overall 242/244 99.2% 97.0–
Influenza A
h
H1
Fresh 0/1 0.0% 0.0–
Frozen 0/0 N/A N/A 1089/1089 100.0% 99.6–
Overall 0/1 0.0% 0.0–
Influenza A
Fresh 62/63 98.4% 91.5–
H1N1/pdm09j
Frozen 18/18 100.0% 82.4–
Overall 80/81 98.8% 93.3–
83.7–
96.8
87.7–
99.0
88.6–
97.2
86.3–
95.1
84.4–
94.4
87.4–
93.9
95.8–
99.9
99.8
99.8
79.3
79.3
99.7
100.0
99.8
828/829 99.9%
99.3–
100.0
1030/1034 99.6%
99.0–
99.8
1858/1863 99.7%
99.4–
99.9
715/739 96.8%
95.2–
97.8
941/953 98.7%
97.8–
99.3
1656/1692 97.9%
97.1–
98.5
753/757 99.5%
98.6–
99.8
972/977 99.5% 98.8–
99.8
1725/1734 99.5% 99.0–
99.7
894/894 100.0% 99.6–
100.0
100.0
1983/1983 100.0% 99.8–
100.0
826/831 99.4% 98.6–
99.7
1071/1071 100.0% 99.6–
100.0
1897/1902 99.7% 99.4–
99.9
(continued on next page)

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
59
Table 9 (continued)
Analyte TP/(TP+FN) Sensitivity/PPA 95% CI TN/(TN+FP) Specificity/NPA 95% CI
Viruses
(continued)
Influenza A
H3j
Influenza Bk Fresh 64/67 95.5%
Parainfluenza
virus 1l
Parainfluenza
virus 2
Parainfluenza
virus 3m
Fresh 67/67 100.0%
Frozen 89/90 98.9%
Overall 156/157 99.4%
Frozen 58/62 93.5%
Overall 122/129 94.6%
Fresh 3/3 100.0%
Frozen 13/14 92.9% 68.5–
Overall 16/17 94.1% 73.0–
Fresh 2/2 100.0% 34.2–
Frozen 0/0 N/A N/A 1089/1089 100.0% 99.6–
Overall 2/2 100.0% 34.2–
Fresh 102/104 98.1% 93.3–
Frozen 9/9 100.0% 70.1–
Overall 111/113 98.2% 93.8–
94.5–
100.0
82.4–
100.0
93.3–
99.8
87.6–
98.5
84.6–
97.5
89.2–
97.3
43.8–
100.0
98.7
99.0
100.0
100.0
99.5
100.0
99.5
825/826 99.9%
992/998 99.4%
1817/1824 99.6%
827/828 99.9%
1026/1026 100.0%
1853/1854 99.9%
892/892 100.0%
1072/1075 99.7% 99.2–
1964/1967 99.8% 99.6–
893/893 100.0% 99.6–
1982/1982 100.0% 99.8–
788/793 99.4% 98.5–
1081/1081 100.0% 99.6–
1869/1874 99.7% 99.4–
(continued on next page)
99.3–
100.0
98.7–
99.7
99.2–
99.8
99.3–
100.0
99.6–
100.0
99.7–
100.0
99.6–
100.0
99.9
99.9
100.0
100.0
100.0
99.7
100.0
99.9

60
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Chlamydophila
Table 9 (continued)
Analyte TP/(TP+FN) Sensitivity/PPA 95% CI TN/(TN+FP) Specificity/NPA 95% CI
Viruses
(continued)
Parainfluenza
virus 4n
Respiratory
Syncytial Virus
o
(RSV)
Bacteria
Bordetella
pertussis
pneumoniae
Mycoplasma
pneumoniae
Fresh 3/3 100.0%
43.8–
892/892 100.0%
100.0
Frozen 0/0 N/A N/A 1087/1089 99.8%
Overall 3/3 100.0%
43.8–
1979/1981 99.9%
100.0
Fresh 73/76 96.1%
88.9–
819/820 99.9%
98.6
Frozen 139/144 96.5%
92.1–
941/945 99.6%
98.5
Overall 212/220 96.4%
93.0–
1760/1765 99.7%
98.1
Fresh 2/2 100.0% 34.2–
p
Frozen 1/1 100.0% 20.7–
100.0
893/893 100.0% 99.6–
1082/1088 99.4% 98.8–
100.0
Overall 3/3 100.0%
43.8–
1975/1981 99.7%
100.0
Fresh 4/4 100.0%
q
Frozen 1/1 100.0%
51.0–
100.0
20.7–
891/891 100.0%
1087/1088 99.9%
100.0
Overall 5/5 100.0%
56.6–
1978/1979 99.9%
100.0
Fresh 18/18 100.0%
r
Frozen 1/1 100.0%
82.4–
100.0
20.7–
875/877 99.8%
1085/1088 99.7%
100.0
Overall 19/19 100.0%
83.2–
1960/1965 99.7%
100.0
99.6–
100.0
99.3–
99.9
99.6–
100.0
99.3–
100.0
98.9–
99.8
99.3–
99.9
100.0
99.7
99.3–
99.9
99.6–
100.0
99.5–
100.0
99.7–
100.0
99.2–
100.0
99.2–
99.9
99.4–
99.9

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
61
a
Adenovirus was detected in 3/4 FN specimens using an independent molecular method. Adenovirus was detected in
6/16 FP specimens using an independent molecular method.
b
The single FN specimen was negative for Coronavirus HKU1 when tested using an independent molecular method.
Coronavirus HKU1 was detected 0/7 FP specimens using an independent molecular method.
c
Coronavirus NL63 was detected in 7/7 FN specimens using an independent molecular method. Coronavirus NL63
was detected in 1/2 FP specimens using an independent molecular method.
d
The 3 FN specimens were negative for Coronavirus OC43 when tested using an independent molecular method.
Coronavirus OC43 was detected in 3/4 FP specimens using an independent molecular method.
e
Human metapneumovirus (hMPV) was detected in 4/7 FN specimens using an independent molecular method.
hMPV was detected in 3/5 FP specimens using an independent molecular method.
f
Rhinovirus was detected in 18/26 FN specimens using an independent molecular method. Rhinovirus was detected
in 14/36 FP specimens using an independent molecular method.
g
Influenza A was detected in 1/2 FN specimens by an independent molecular method. Three (3) FP samples were not
available for testing. Influenza A was detected in the 3/6 remaining FP samples by an independent molecular
method.
h
Influenza A H1 was detected in 1/1 FN specimen by an independent molecular method. Note: Non-2009 H1 has
not been in circulation since being replaced by the 2009 H1 and thus this discrepancy test result is likely false.
i
Influenza A H1N1 pdm09 was detected in 1/1 FN by an independent molecular method. Influenza A H1 was
detected in 3/5 FP specimens by an independent molecular method.
j
Influenza A H3 was detected in 1/1 FN by an independent molecular method. Influenza H3 was detected in 7/7 FP
specimens by an independent molecular method.
k
Influenza B was detected in 6/6 FN specimens available for testing by an independent molecular method; one
discordant sample was not tested by an independent molecular method. Influenza B was detected in 1/1 FP
specimens available for testing by an independent molecular method.
l
The single FN specimen was negative for Parainfluenza virus 1 by an independent molecular method. Parainfluenza
virus 1 was detected in 3/3 FP specimens by an independent molecular method.
m
Parainfluenza virus 3 was detected in 1/2 FN specimens by an independent molecular method. Parainfluenza 3 was
detected in 3/5 FP specimens by an independent molecular method.
n
Parainfluenza virus 4 was detected in 2/2 FP specimens by an independent molecular method.
o
Respiratory Syncytial Virus was detected in 2/8 FN specimens by an independent molecular method. Respiratory
Syncytial Virus was detected in 3/5 FP specimens by an independent molecular method.
p
Bordetella pertussis
q
Chlamydophila pneumoniae
r
Mycoplasma pneumoniae
was detected in 1/6 FP specimens by an independent molecular method.
was detected in 1/1 FP specimens by an independent molecular method.
was detected in 1/4 specimens by an independent molecular method.
The QIAstat-Dx Respiratory Panel detected a total of 191 specimens with distinctive multiple
organism detections (9.6% of all specimens) in the prospective study.
All distinct co-infection combinations, as detected by the QIAstat-Dx Respiratory Panel during
the prospective clinical study, are presented in Table 10
(next page).

62
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Table 10. Distinct co-infection combinations detected by QIAstat-Dx Respiratory Panel in the prospective study
Distinct co-infection combinations detected by the QIAstat-Dx
Respiratory Panel
Analyte 1 Analyte 2 Analyte 3 Analyte 4
Adenovirus
Adenovirus
Adenovirus
Adenovirus
Rhinovirus/
Enterovirus
Rhinovirus/
Enterovirus
Respiratory
Syncytial Virus
Mycoplasma
Coronavirus
NL63
12 3
11 1
2 1
2 0 N/A
Total coinfections
pneumoniae
Adenovirus
Coronavirus
HKU1
Coronavirus
HKU1
Coronavirus
HKU1
Coronavirus
HKU1
Coronavirus
HKU1
Coronavirus
HKU1
Coronavirus
HKU1
Coronavirus
NL63
Coronavirus
NL63
Coronavirus
NL63
Coronavirus
HKU1
Adenovirus Respiratory
Human
Metapneumovirus
Parainfluenza
virus 3
Parainfluenza
virus 4
Respiratory
Syncytial Virus
Rhinovirus/
Enterovirus
Rhinovirus/
Enterovirus
Adenovirus
Adenovirus 1 1 Adenovirus (1)
Bordetella
3 1
Syncytial
Virus
3 1
Rhinovirus/
Enterovirus
1 1
8 1
Respiratory
Syncytial
Virus
4 1
Respiratory
Syncytial
Virus
2 2
1 1 Coronavirus HKU1
1 0 N/A
1 0 N/A
1 0 N/A
pertussis
Number of
discrepant
co-infections
Discrepant analyte(s)
Rhinovirus/Enterovirus
(1); Adenovirus (2)
Respiratory Syncytial
Virus (1)
Mycoplasma
pneumoniae
Coronavirus HKU1
(1)
(1)
Human
Metapneumovirus (1)
Coronavirus HKU1,
Parainfluenza virus 4
(1)
Coronavirus HKU1
(1)
Rhinovirus/Enterovirus
(1)
Bordetella pertussis
(2)
(continued on next page)
(1)

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
63
Table 10 (continued)
Distinct co-infection combinations detected by the QIAstat-Dx
Respiratory Panel
Analyte 1 Analyte 2 Analyte 3 Analyte 4
Coronavirus
NL63
Coronavirus
NL63
Coronavirus
NL63
Coronavirus
OC43
Coronavirus
OC43
Coronavirus
OC43
Coronavirus
OC43
Coronavirus
OC43
Coronavirus
OC43
Coronavirus
229E
Human
Metapneumovirus
Human
Metapneumovirus
Human
Metapneumovirus
Human
Metapneumovirus
Influenza A (no
subtype)
Parainfluenza
virus 1
Respiratory
Syncytial Virus
Rhinovirus/
Enterovirus
Adenovirus
Human
Metapneumovirus
Parainfluenza
virus 3
Respiratory
Syncytial Virus
Rhinovirus/
Enterovirus
Rhinovirus/
Enterovirus
Respiratory
Syncytial Virus
Adenovirus 2 1 Adenovirus (1)
Respiratory
Syncytial Virus
Rhinovirus/
Enterovirus
Rhinovirus/
Enterovirus
Respiratory
Syncytial Virus
1 0 N/A
2 0 N/A
2 0 N/A
2 0 N/A
2 0 N/A
Rhinovirus/
Enterovirus
4 0 N/A
Respiratory
Syncytial
Virus
2 2 Rhinovirus/
1 0 N/A
2 0 N/A
9 3 Rhinovirus/
Adenovirus Coronavirus
Adenovirus 1 1 Influenza A,
1 0 N/A
2 0 N/A
229E
Total coinfections
1 1 Adenovirus,
Number of
discrepant
coinfections
Discrepant
analyte(s)
Enterovirus (2)
Enterovirus (3)
Rhinovirus/
Enterovirus (1)
Adenovirus (1)
(continued on next page)

64
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Table 10 (continued)
Distinct co-infection combinations detected by the QIAstat-Dx
Respiratory Panel
Analyte 1 Analyte 2 Analyte 3 Analyte 4
Influenza A (no
subtype)
Influenza A
H1N1/pdm09
Influenza A
H1N1/pdm09
Influenza A
H1N1/pdm09
Influenza A
H1N1/pdm09
Influenza A
H1N1/pdm09
Influenza A H3 Adenovirus 2 1 Adenovirus (1)
Influenza A H3 Coronavirus NL63 Parainfluenza
Influenza A H3 Coronavirus NL63
Respiratory
Syncytial Virus
Coronavirus NL63 1 0 N/A
Coronavirus
OC43
Rhinovirus/
Enterovirus
Rhinovirus/
Enterovirus
Respiratory
Syncytial Virus
1 0 N/A
Adenovirus 1 1 Adenovirus (1)
2 0 N/A
Bordetella
1 0 N/A
pertussis
1 0 N/A
virus 1
Bordetella
1 0 N/A
1 1
Total coinfections
pertussis
Influenza A H3 Coronavirus NL63 1 1 NL63 (1)
Influenza A H3
Influenza A H3
Influenza A H3
Influenza A H3
Influenza A H3
Influenza A H3 Coronavirus 229E 1 0 N/A
Influenza B Coronavirus HKU1 3 0 N/A
Influenza B Coronavirus NL63 1 0 N/A
Coronavirus
OC43
Rhinovirus/
Enterovirus
Parainfluenza
virus 1
Parainfluenza
virus 3
Respiratory
Syncytial Virus
Adenovirus
4 2
2 0 N/A
2 0 N/A
1 0 N/A
Respiratory
Syncytial
Virus
1 1
Number of
discrepant
co-infections
(continued on next page)
Discrepant
analyte(s)
Bordetella
pertussis
(1)
Coronavirus
OC43,
Adenovirus (1)
Rhinovirus/
Enterovirus (2)

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
65
Mycoplasma
Table 10 (continued)
Distinct co-infection combinations detected by the QIAstat-Dx
Respiratory Panel
Analyte 1 Analyte 2 Analyte 3 Analyte 4
Influenza B Respiratory
Influenza B Rhinovirus/
Mycoplasma
pneumoniae
Mycoplasma
pneumoniae
Parainfluenza
virus 1
Parainfluenza
virus 1
Parainfluenza
virus 1
Parainfluenza
virus 1
Parainfluenza
virus 3
Parainfluenza
virus 3
Parainfluenza
virus 3
Parainfluenza
virus 3
Parainfluenza
virus 3
Parainfluenza
virus 4
Parainfluenza
virus 4
Syncytial Virus
Enterovirus
Coronavirus
HKU1
Rhinovirus/
Enterovirus
Adenovirus 1 0 N/A
Respiratory
Syncytial Virus
Rhinovirus/
Enterovirus
Rhinovirus/
Enterovirus
Adenovirus 3 2 Adenovirus (2)
Adenovirus Rhinovirus/
Human
Metapneumovirus
Respiratory
Syncytial Virus
Rhinovirus/
Enterovirus
Respiratory
Syncytial Virus
Rhinovirus/
Enterovirus
Number of
Total coinfections
2 0 N/A
7 4 Rhinovirus/
1 1 Coronavirus HKU1
1 0 N/A
discrepant
coinfections
1 1 Parainfluenza virus 1
2 0 N/A
1 1 Rhinovirus/
pneumoniae
Enterovirus
2 1
2 1
14 3
1 0 N/A
2 0 N/A
3 1 Parainfluenza virus 3
(continued on next page)
Discrepant analyte(s)
Enterovirus (4)
(1)
(1)
Enterovirus (1)
(1)
Human
Metapneumovirus (1)
Parainfluenza virus 3
(1)
Rhinovirus/
Enterovirus (2),
Parainfluenza virus 3
(1)

66
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Table 10 (continued)
Distinct co-infection combinations detected by the QIAstat-Dx
Respiratory Panel
Analyte 1 Analyte 2 Analyte 3 Analyte 4
Respiratory
Syncytial
Virus
Respiratory
Syncytial
Virus
Respiratory
Syncytial
Virus
Rhinovirus/
Enterovirus
Total co-infections 191 51
Total double infections 166 42
Total triple infections 22 7
Total quadruple infections 3 2
The three organisms most prevalent in multiple detections by the QIAstat-Dx Respiratory Panel
Human
Metapneumovirus
Human
Metapneumovirus
Rhinovirus/
Enterovirus
Respiratory
Syncytial Virus
Rhinovirus/
Enterovirus
Rhinovirus/
Enterovirus
29 6 Rhinovirus/
Adenovirus 2 0 N/A
Adenovirus 1 0 N/A
2 1
Total coinfections
Number of
discrepant
coinfections
Discrepant analyte(s)
Human
Metapneumovirus,
Rhinovirus/
Enterovirus (1)
Enterovirus (5),
Respiratory Syncytial
Virus (1)
in the prospective study were Rhinovirus/Enterovirus (108/191, 56.5%), Respiratory Syncytial
Virus (77/191, 40.8%), and Adenovirus (53/191, 27.7%). The prevalence of individual
organisms in each multiple detection are shown in Table 11 (next page).

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
67
Table 11. The prevalence of individual organisms in each QIAstat-Dx multiple detection
Analyte Prevalence in multiple detections (N=191)
Viruses
Adenovirus 53 (27.7%)
Coronavirus 229E 3 (1.6%)
Coronavirus HKU1 26 (13.6%)
Coronavirus NL63 16 (8.4%)
Coronavirus OC43 15 (7.9%)
Human Metapneumovirus 24 (12.6%)
Rhinovirus/Enterovirus 108 (56.5%)
Influenza A H1 0 (0.0%)
Influenza A H1N1/pdm09 6 (3.1%)
Influenza A H3 16 (8.4%)
Influenza B 13 (6.8%)
Parainfluenza virus 1 9 (4.7%)
Parainfluenza virus 2 0 (0.0%)
Parainfluenza virus 3 28 (14.7%)
Parainfluenza virus 4 4 (2.1%)
Respiratory Syncytial Virus 78 (40.8%)
Bacteria
Bordetella pertussis
Chlamydophila pneumoniae
Mycoplasma pneumoniae
4 (2.1%)
0 (0.0%)
5 (2.6%)
Additional distinct co-infection combinations detected by the comparator method but not
detected by the QIAstat-Dx Respiratory Panel in the prospective clinical trial are presented in
Table 12 (next page).

68
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Table 12. Additional distinct co-infection combinations detected by the comparator method but not by the QIAstat-Dx
Respiratory Panel in the prospective study
Distinct co-infection combinations detected by the comparator method
Analyte 1 Analyte 2 Analyte 3
Adenovirus Coronavirus HKU1 Respiratory Syncytial Virus 1
Adenovirus Coronavirus OC43 Coronavirus NL63 1
Adenovirus Respiratory Syncytial Virus Coronavirus NL63 1
Adenovirus Rhinovirus/Enterovirus Respiratory Syncytial Virus 1
Coronavirus HKU1 Coronavirus OC43 1
Coronavirus HKU1 Respiratory Syncytial Virus 1
Coronavirus HKU1 Coronavirus NL63 Respiratory Syncytial Virus 1
Coronavirus HKU1 Coronavirus NL63 1
Coronavirus HKU1 Parainfluenza virus 1 Rhinovirus/Enterovirus 1
Coronavirus NL63 Respiratory Syncytial Virus 1
Coronavirus NL63 Rhinovirus/Enterovirus 1
Coronavirus NL63 Influenza A H3 1
Coronavirus OC43 Respiratory Syncytial Virus 1
Human Metapneumovirus Parainfluenza virus 3 Rhinovirus/Enterovirus 1
Human Metapneumovirus Rhinovirus/Enterovirus 1
Rhinovirus/Enterovirus Adenovirus 1
Rhinovirus/Enterovirus Influenza A H3 2
Rhinovirus/Enterovirus Parainfluenza virus 3 1
Rhinovirus/Enterovirus Parainfluenza virus 3 Respiratory Syncytial Virus 1
Rhinovirus/Enterovirus Parainfluenza virus 4 2
Influenza A H3 Respiratory Syncytial Virus 1
Influenza B Influenza A (Equivocal) 1
Total co-infections 24
Total double infections 16
Total triple infections 8
Total co-infections

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
69
A total of 1994 prospective clinical specimens were tested and analyzed during the
prospective clinical evaluation. Of these, 95.88% (1912/1994) yielded valid results on the
first attempt (i.e., first loaded cartridge). Invalid or no result were obtained for the remaining
82 specimens (4.11%). Forty-two (42) specimens were invalid due to cartridge internal control
failure (2.11%). Of these, 20 (1.00%) provided a result for positively detected targets and
22 (1.10%) had no detections. For 40 (2.00%) specimens, no results were obtained due to
incomplete runs. Of these, 1 specimen was aborted by users (0.05%), 21 were due to
instrument errors (1.05%), and 18 were due to cartridge-related errors (0.90%). Seventy-two
(72) of the 82 initially failed (no result or invalid); specimens yielded valid results after a single
retesting using a new cartridge/sample. The remaining 10 specimens failed on the second
attempt (2 due to cartridge failures, 1 due to instrument errors, and 7 due to internal control
failures). Of these internal control failures, detected pathogens were reported for 4 specimens.
Preselected archived specimens
Some of the analytes on the QIAstat-Dx Respiratory Panel were of low prevalence and were
not encountered in sufficiently large numbers during the prospective study to adequately
demonstrate clinical performance. To supplement the results of the prospective clinical study,
an evaluation of preselected frozen archived retrospective specimens was performed. The
specimens selected for testing had previously tested positive for one of the following targets at
the clinical laboratory by their standard of care method:
Bordetella pertussis
229E, Coronavirus HKU1, Coronavirus NL63, Coronavirus OC43, Influenza A H1N1 2009,
Mycoplasma pneumoniae, Chlamydophila pneumoniae
, Parainfluenza virus 1, Parainfluenza
virus 2, and Parainfluenza virus 4. Testing was performed by operators who were blinded to
the expected test result. A total of 310 clinical samples were included within the frozen
archived retrospective sample tested arm. Samples were tested by both the comparator method
and QIAstat-Dx Respiratory Panel. If the comparator method did not confirm the preselected
target as positive, it was excluded from the data analysis for that target.
, Coronavirus
A summary of the demographic information available for the archived specimens is provided
in Table 13 (next page).

70
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Table 13. Demographic summary for the retrospective study arm
Overall (%)
Male 158 (50.8%)
SEX
Female 152 (49.2%)
≤5 years 139 (44.9%)
6–21 years 85 (27.4%)
22–49 years 53 (17.1%)
AGE
50+ years 33 (10.7%)
Outpatient 224 (72.3%)
Hospitalized 68 (21.9%)
Emergency 8 (2.6%)
ICU 8 (2.6%)
STATUS
Other 2 (0.6%)
Total 310
The QIAstat-Dx Respiratory Panel retrospective specimens testing performance data against
the comparator method are provided in Table 14 (next page) by analyte.

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
71
Table 14. Overall retrospective clinical study performance
Analyte TP/(TP+FN) Sensitivity
/PPA
Viruses
95% CI TN/(TN+FP) Specificity
/NPA
95% CI
Adenovirusa 9/9 100.0% 70.1–100.0 297/304 97.8% 95.4–98.9
Coronavirus 229E 26/27 96.3% 81.7–99.3 286/286 100.0% 98.7–100.0
Coronavirus HKU1b 14/14 100.0% 78.5–100.0 298/299 99.7% 98.1–99.9
Coronavirus NL63c 24/24 100.0% 86.2–100.0 286/288 99.3% 97.5–99.8
Coronavirus OC43 28/29 96.6% 82.8–99.4 279/279 100.0% 98.6–100.0
Human Metapneumovirus 2/2 100.0% 34.2–100.0 311/311 100.0% 98.7–100.0
Rhinovirus/
44/49 89.8% 78.2–95.5 254/264 96.2% 93.2–97.9
Enterovirusd
Influenza A 17/17 100.0% 81.5–100.0 296/296 100.0% 98.7–100.0
Influenza A H1 0/0 N/A N/A 313/313 100.0% 98.8–100.0
Influenza A
H1N1/pdm09
e
7/8 87.5% 52.9–97.8 304/304 100.0% 98.8–100.0
Influenza A H3 8/8 100.0% 67.5–100.0 305/305 100.0% 98.8–100.0
Influenza B 1/1 100.0% 20.7–100.0 312/312 100.0% 98.8–100.0
Parainfluenza virus 1 40/40 100.0% 91.2–100.0 267/267 100.0% 98.6–100.0
Parainfluenza virus 2 3/3 100.0% 43.8–100.0 309/309 100.0% 98.8–100.0
Parainfluenza virus 3f 1/4 25.0% 4.6–69.9 309/309 100.0% 98.8–100.0
Parainfluenza virus 4g 22/24 91.7% 74.2–97.7 278/278 100.0% 98.6–100.0
Respiratory Syncytial
11/12 91.7% 64.6–98.5 300/301 99.7% 98.4–99.9
Virus (RSV)h
Bacteria
Bordetella pertussis
Chlamydophila
pneumoniae
i
Mycoplasma
33/33 100.0% 89.6–100.0 261/261 100.0% 98.5–100.0
54/61 88.5% 78.2–94.3 250/250 100.0% 98.5–100.0
25/25 100.0% 86.7–100.0 287/288 99.7% 98.1–99.9
pneumoniae
a
Adenovirus was detected in 3/5 FP specimens using an independent molecular method. 2 FP did not undergo
discordant analysis.
b
The single FP Coronavirus HKU1 specimen was negative when tested using an independent molecular method.
c
The single FP Coronavirus NL63 specimen was negative when tested using an independent molecular method.

72
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
d
Rhinovirus was detected in 1/2 FN when tested using an independent molecular method. Rhinovirus was detected in
4/10 FP specimens using an independent molecular method.
e
Influenza H1N1 pdm09 was detected in the single FN specimen.
f
Parainfluenza virus 3 was detected in 1/3 FN specimens by an independent molecular method.
g
Parainfluenza virus 4 was detected in 1/2 FN specimens by an independent molecular method.
h
The single FN Respiratory Syncytial Virus was negative for that target by an independent molecular method. The
single FP Respiratory Syncytial Virus was negative for that target by an independent molecular method.
i
Chlamydophila pneumoniae
was detected in 4/5 FN specimens by an independent molecular method.
Testing of contrived specimen
Influenza A H1, Parainfluenza virus 2, Parainfluenza virus 4, Coronavirus 229E and
Chlamydophila pneumoniae
insufficient to demonstrate system performance. Therefore, contrived specimens were used as
surrogate clinical specimens to supplement and test the sensitivity and specificity of the above
analytes. Residual negative clinical specimens were spiked with the pathogens at 3x, 5x and
10x LoD levels (50 of each).
Contrived samples were provided a unique study identification number and the individual who
contrived the samples did not test them therefore the status of each contrived specimen was
unknown at the time of testing. Results of contrived specimen testing are provided in Table 15
(next page).
, despite all prospective and retrospective testing efforts, were

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
73
Table 15. Contrived specimen results
Positive Predictive
Agreement
Analyte
Influenza A H1* 3 24/24 100% 86.2–100
Coronavirus 229E 3 16/16 100% 80.6–100
Parainfluenza virus 2 3 16/16 100% 80.6–100
Parainfluenza virus 4 3 15/16 93.8% 71.7–98.9
Chlamydophila
pneumoniae
* One Influenza A H1 strain [VR-897] was initially spiked incorrectly, yielding unexpected results across all LoD
concentrations [3x LoD = 4/8 (50%), 5x LoD = 2/9 (22.2%) and 10x LoD = 6/8 (75.0%). A replacement strain
[0810244CFHI] was sent to the testing site for spiking and strain VR-897 was also repeated to confirm that the issue
was isolated to a procedural error and not an instrument failure.
x LoD TP/(TP + FN) % 95% CI
5 27/27 100% 87.5–100
10 24/24 100% 86.2–100
5 18/18 100% 82.4–100
10 16/16 100% 80.6–100
5 18/18 100% 82.4–100
10 16/16 100% 80.6–100
5 18/18 100% 82.4–100
10 16/16 100% 80.6–100
3 16/16 100% 80.6–100
5 18/18 100% 82.4–100
10 16/16 100% 80.6–100
Expected values
During the prospective QIAstat-Dx Respiratory Panel (Cat. No. 691221) clinical study,
1994 eligible prospective nasopharyngeal swab (NPS) specimens were collected and tested
at five (5) sites across the U.S. (4) and Europe (1) from December 2017 through June 2018.
The number and percentage of positive cases, as determined by the QIAstat-Dx Respiratory
Panel, calculated by testing site or by age group are presented in Table 16, Table 17, and
Table 18 (following pages).

74
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Table 16. Expected value (EV) (as determined by the QIAstat Dx Respiratory Panel) summary overall and by site for the
prospective clinical evaluation (N = number)
Overall
Organism N EV N EV N EV N EV N EV N EV
Viruses
Adenovirus 102 5.1% 44 10.5% 9 2.1% 12 2.7% 30 9.9% 7 1.8%
Coronavirus 229E 8 0.4% 1 0.2% 0 0.0% 0 0.0% 7 2.3% 0 0.0%
Coronavirus HKU1 58 2.9% 4 1.0% 11 2.6% 14 3.1% 12 4.0% 17 4.3%
Coronavirus NL63 42 2.1% 4 1.0% 1 0.2% 15 3.3% 11 3.6% 11 2.8%
Coronavirus OC43 30 1.5% 0 0.0% 5 1.2% 6 1.3% 12 4.0% 7 1.8%
Human
Metapneumovirus
Human
Rhinovirus/Enterovirus
Influenza A 251 12.6% 120 28.7% 0 0.0% 58 12.9% 38 12.5% 35 8.8%
Influenza A H1 0 0.0% 0 0.0% 0 0.0% 0 0.0% 0 0.0% 0 0.0%
Influenza A H1N1
pdm 2009
Influenza H3 163 8.2% 52 12.4% 0 0.0% 52 11.6% 28 9.2% 31 7.8%
Influenza B 123 6.2% 58 13.9% 0 0.0% 32 7.1% 7 2.3% 26 6.5%
Parainfluenza virus 1 19 1.0% 2 0.5% 1 0.2% 2 0.4% 4 1.3% 10 2.5%
Parainfluenza virus 2 2 0.1% 2 0.5% 0 0.0% 0 0.0% 0 0.0% 0 0.0%
Parainfluenza virus 3 116 5.8% 23 5.5% 19 4.5% 16 3.6% 23 7.6% 35 8.8%
Parainfluenza virus 4 5 0.3% 1 0.2% 0 0.0% 1 0.2% 0 0.0% 3 0.8%
Respiratory Syncytial
Virus
Bacteria
Bordetella pertussis
Chlamydophila
pneumoniae
Mycoplasma
pneumoniae
(n=1994) Site 1 (n=418) Site 2 (n=426) Site 3 (n=448) Site 4 (n=303) Site 5 (n=399)
120 6.0% 42 10.0% 24 5.6% 14 3.1% 14 4.6% 26 6.5%
304 15.2% 59 14.1% 78 18.3% 39 8.7% 53 17.5% 75 18.8%
85 4.3% 67 16.0% 0 0.0% 4 0.9% 10 3.3% 4 1.0%
217 10.9% 64 15.3% 40 9.4% 35 7.8% 40 13.2% 38 9.5%
9 0.5% 2 0.5% 1 0.2% 0 0.0% 6 2.0% 0 0.0%
6 0.3% 2 0.5% 1 0.2% 1 0.2% 1 0.3% 1 0.3%
24 1.2% 19 4.5% 0 0.0% 2 0.4% 1 0.3% 2 0.5%

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
75
Table 17. Expected value (EV) (as determined by the QIAstat Dx Respiratory Panel) summary by age category for the
prospective clinical evaluation (N = number)
Organism Overall (n=1994)
N EV N EV N EV N EV N EV
Viruses
Adenovirus 102 5.1% 78 12.4% 7 2.9% 11 3.3% 6 0.8%
Coronavirus 229E 8 0.4% 4 0.6% 4 1.7% 0 0.0% 0 0.0%
Coronavirus HKU1 58 2.9% 29 4.6% 5 2.1% 8 2.4% 16 2.0%
Coronavirus NL63 42 2.1% 25 4.0% 3 1.3% 5 1.5% 9 1.1%
Coronavirus OC43 30 1.5% 20 3.2% 2 0.8% 4 1.2% 4 0.5%
Human
Metapneumovirus
Human
Rhinovirus/Enterovirus
Influenza A 251 12.6% 47 7.5% 36 15.1% 64 19.4% 104 13.0%
Influenza A H1 0 0.0% 0 0.0% 0 0.0% 0 0.0% 0 0.0%
Influenza A H1N1
pdm 2009
Influenza H3 163 8.2% 25 4.0% 30 12.6% 35 10.6% 73 9.1%
Influenza B 123 6.2% 11 1.8% 22 9.2% 27 8.2% 63 7.9%
Parainfluenza virus 1 19 1.0% 11 1.8% 0 0.0% 4 1.2% 4 0.5%
Parainfluenza virus 2 2 0.1% 1 0.2% 0 0.0% 0 0.0% 1 0.1%
Parainfluenza virus 3 116 5.8% 70 11.2% 4 1.7% 6 1.8% 36 4.5%
Parainfluenza virus 4 5 0.3% 4 0.6% 0 0.0% 0 0.0% 1 0.1%
Respiratory Syncytial
Virus
Bacteria
Bordetella pertussis
Chlamydophila
pneumoniae
Mycoplasma
pneumoniae
120 6.0% 46 7.3% 3 1.3% 17 5.2% 54 6.8%
304 15.2% 186 29.7% 35 14.6% 22 6.7% 61 7.6%
85 4.3% 20 3.2% 6 2.5% 30 9.1% 29 3.6%
217 10.9% 135 21.5% 11 4.6% 17 5.2% 54 6.8%
9 0.5% 5 0.8% 2 0.8% 0 0.0% 2 0.3%
6 0.3% 1 0.2% 3 1.3% 2 0.6% 0 0.0%
24 1.2% 4 0.6% 6 2.5% 11 3.3% 3 0.4%
≤5 years
(n=627)
6–21 years
(n=239)
22–49 years
(n=330)
>49 years
(n=798)
The number and percentage of co-infection cases, as determined by the QIAstat-Dx Respiratory
Panel, calculated by age group are presented in Table 17,next page.

76
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Table 18. Expected value (co-infections as determined by the QIAstat-Dx Respiratory Panel) summary by age group
Number
Co-infection
AdV + HRV/EV
+ CoV NL63
AdV + HRV/EV 12 (6.28%) 9 2 1 0
AdV + RSV 11 (5.82%) 11 0 0 0
AdV +
M.
pneumoniae
AdV + CoV
HKU1
CoV HKU1+
AdV+ RSV
CoV HKU1 +
HMPV
CoV HKU1 +
PIV 3 + HRV/EV
CoV HKU1 +
PIV 4
CoV HKU1 +
RSV
CoV HKU1 +
HRV/EV + RSV
CoV HKU1 +
HRV/EV
CoV NL63 +
AdV+ RSV
CoV NL63 +
AdV
CoV NL63 +
pertussis
(expected value)
overall (n=191)
2 (1.05%) 2 0 0 0
2 (1.05%) 1 1 0 0
3 (1.57%) 3 0 0 0
1 (0.52%) 1 0 0 0
3 (1.57%) 3 0 0 0
1 (0.52%) 1 0 0 0
1 (0.52%) 1 0 0 0
8 (4.28%) 5 1 1 1
1 (0.52%) 1 0 0 0
4 (2.09%) 2 0 0 2
1 (0.52%) 1 0 0 0
1 (0.52%) 1 0 0 0
B.
2 (1.05%) 1 1 0 0
<6 years
(n=151)
6–21 years
(n=12)
22–49 years
(n=14)
>49 years
(n=14)
(continued on next page)

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
77
Table 18 (continued)
Number
Co-infection
CoV NL63 + PIV 1 1 (0.52%) 0 0 1 0
(expected value)
overall (n=191)
<6 years
(n=151)
6–21 years
(n=12)
22–49 years
(n=14)
>49 years
(n=14)
CoV NL63 +
RSV
CoV NL63 +
HRV/EV
CoV OC43 +
AdV
CoV OC43 +
HMPV
CoV OC43 +
PIV 3 + HRV/EV
CoV OC43 +
RSV
CoV OC43 +
HRV/EV + RSV
CoV OC43 +
HRV/EV
CoV 229E +
RSV
HMPV + AdV 2 (1.05%) 1 0 1 0
HMPV + RSV 2 (1.05%) 1 0 0 1
HMPV +
HRV/EV
HMPV +
HRV/EV + AdV
+ CoV 229E
Influenza A (no
subtype) + RSV
+ AdV
Influenza A (no
subtype) + RSV
2 (1.05%) 2 0 0 0
2 (1.05%) 2 0 0 0
2 (1.05%) 2 0 0 0
2 (1.05%) 2 0 0 0
1 (0.52%) 1 0 0 0
4 (2.09%) 3 1 0 0
2 (1.05%) 2 0 0 0
2 (1.05%) 1 1 0 0
1 (0.52%) 1 0 0 0
9 (4.71%) 6 0 2 1
1 (0.52%) 1 0 0 0
1 (0.52%) 1 0 0 0
1 (0.52%) 1 0 0 0
(continued on next page)

78
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Table 18 (continued)
Co-infection
Influenza A
H1N1/pdm09
+ CoV NL63
Influenza A
H1N1/pdm09
+ CoV OC43 +
AdV
Influenza A
H1N1/pdm09
+ HRV/EV
Influenza A
H1N1/pdm09
+ HRV/EV +
B. pertussis
Influenza A
H1N1/pdm09
+ RSV
Influenza A H3
+ AdV
Influenza A H3
+ CoV NL63 +
PIV 1
Influenza A H3
+ CoV NL63 +
B. pertussis
Influenza A H3
+ CoV NL63
Influenza A H3
+ CoV OC43 +
AdV + RSV
Influenza A H3
+ HRV/EV
Influenza A H3
+ PIV 1
Influenza A H3
+ PIV 3
Number
(expected value)
overall (n=191)
1 (0.52%) 0 0 1 0
1 (0.52%) 1 0 0 0
2 (1.05%) 1 1 0 0
1 (0.52%) 1 0 0 0
1 (0.52%) 1 0 0 0
2 (1.05%) 0 0 1 1
1 (0.52%) 1 0 0 0
1 (0.52%) 1 0 0 0
1 (0.52%) 0 0 0 1
1 (0.52%) 1 0 0 0
4 (2.09%) 2 0 1 1
2 (1.05%) 2 0 0 0
2 (1.05%) 1 0 0 1
<6 years
(n=151)
6–21 years
(n=12)
22–49 years
(n=14)
(continued on next page)
>49 years
(n=14)

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
79
Table 18 (continued)
Number
Co-infection
Influenza A H3
+ RSV
Influenza A H3
+ CoV 229E
Influenza B +
CoV HKU1
Influenza B +
CoV NL63
Influenza B +
RSV
Influenza B +
HRV/EV
M. pneumoniae
+ CoV HKU1
M. pneumoniae
+ HRV/EV
PIV 1 + AdV 1 (0.52%) 1 0 0 0
PIV 1 + RSV 1 (0.52%) 1 0 0 0
PIV 1 + HRV/EV 2 (1.05%) 2 0 0 0
PIV 1 + HRV/EV
+
M.
pneumoniae
PIV 3 + AdV 3 (1.57%) 3 0 0 0
PIV 3 + AdV +
HRV/EV
PIV 3 + HMPV 2 (1.05%) 2 0 0 0
PIV 3 + RSV 2 (1.05%) 2 0 0 0
PIV 3 + HRV/EV 14 (7.33%) 14 0 0 0
PIV 4 + RSV 1 (0.52%) 1 0 0 0
PIV 4 + HRV/EV 2 (1.05%) 2 0 0 0
(expected value)
overall (n=191)
1 (0.52%) 0 0 1 0
1 (0.52%) 0 1 0 0
3 (1.57%) 1 0 0 2
1 (0.52%) 0 1 0 0
2 (1.05%) 2 0 0 0
7 (3.67%) 4 1 1 1
1 (0.52%) 0 1 0 0
1 (0.52%) 0 0 1 0
1 (0.52%) 1 0 0 0
3 (1.57%) 3 0 0 0
<6 years
(n=151)
6–21 years
(n=12)
22–49 years
(n=14)
>49 years
(n=14)
(continued on next page)

80
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Table 18 (continued)
Number
Co-infection
RSV + HMPV +
HRV/EV + AdV
RSV + HMPV +
HRV/EV
RSV + HRV/EV 29 (15.18%) 26 0 2 1
HRV/EV + RSV
+ AdV
(expected value)
overall (n=191)
1 (0.52%) 1 0 0 0
2 (1.05%) 1 0 0 1
2 (1.05%) 2 0 0 0
<6 years
(n=151)
6–21 years
(n=12)
22–49 years
(n=14)
>49 years
(n=14)
Analytical performance
Sensitivity (Limit of Detection)
The Analytical Sensitivity, or Limit of Detection (LoD), is defined as the lowest concentration at
which ≥95% of the tested samples generate a positive call.
The LoD for each QIAstat-Dx Respiratory Panel pathogen was assessed by analyzing serial
dilutions of analytical samples prepared from high-titer stocks obtained from commercial
®
suppliers (ZeptoMetrix and ATCC
analytes.
) or artificial samples for commercially unavailable target
The LoD concentration was determined for a total of 51 pathogen strains. The LoD per analyte
was determined using selected strains representing individual pathogens that are possible to
detect with the QIAstat-Dx Respiratory Panel. To confirm the established LoD concentration,
the detection rate of all replicates must be ≥95% (at least 19/20 replicates must generate a
positive signal).
At least three different cartridge lots and at least three different QIAstat-Dx Analyzers were
used for LoD determination for every pathogen.
Individual LoD values for each QIAstat-Dx Respiratory Panel target is shown in Table 19 (next page).

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
81
Table 19. LoD values obtained for the different respiratory target strains tested with the QIAstat-Dx Respiratory
SARS-CoV-2 Panel
Pathogen Strain Source Concentration Detection rate
Influenza A H1N1
Influenza A H3N2 A/Virginia/ATCC6/2012* ATCC VR-1811 0.1 PFU/ml Flu A: 20/20
Influenza A, subtype
H1N1/2009/pdm09
Influenza B B/Virginia/ATCC5/2012* ATCC VR-1807 0.03 PFU/ml 20/20
Coronavirus 229E – ATCC VR-740 0.2 TCID50/ml 20/20
Coronavirus OC43 – ATCC VR-1558 0.1 TCID50/ml 20/20
Coronavirus NL63 –
A/New Jersey/8/76 ATCC VR-897 341 CEID
A/Brisbane/59/07
A/New Caledonia/20/99 ZeptoMetrix
A/Wisconsin/67/2005*
A/Port Chalmers/1/73 ATCC VR-810 499.3 CEID50/ml Flu A: 20/20
A/Virginia/ATCC1/2009 ATCC VR-1736 67 PFU/ml Flu A: 20/20
A/SwineNY/03/2009 ZeptoMetrix
B/FL/04/06 ATCC VR-1804 1080 CEID50/ml 20/20
B/Taiwan/2/62 ATCC VR-295 5000 CEID50/ml 19/20
–* ZeptoMetrix
–*
ZeptoMetrix
0810244CFHI
0810036CFHI
ZeptoMetrix
0810252CFHI
0810249CFHI
0810229CFHI
ZeptoMetrix
0810024CFHI
ZeptoMetrix
0810228CFHI
4 TCID50/ml Flu A: 20/20
15 TCID50/ml Flu A: 20/20
3.8 TCID50/ml Flu A: 20/20
56 TCID50/ml Flu A: 20/20
3.6 TCID50/ml 20/20
0.1 TCID50/ml 20/20
0.01 TCID50/ml 20/20
/ml Flu A: 20/20
50
H1: 20/20
H1: 20/20
H1: 19/20
H3: 20/20
H3: 20/20
H3: 20/20
H1N1: 20/20
H1N1: 20/20
Coronavirus HKU1 –* Clinical sample
S510
40,000 copies/ml 20/20
(continued on next page)

82
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Table 18 (continued)
Pathogen Strain Source Concentration Detection rate
Parainfluenza virus 1
(PIV 1)
Parainfluenza virus 2
(PIV 2)
Parainfluenza virus 3
(PIV 3)
Parainfluenza virus
4a (PIV 4a)
Parainfluenza virus
4b (PIV 4b)
Respiratory Syncytial
Virus A
Respiratory Syncytial
Virus B
Human
Metapneumovirus
Adenovirus GB (Adenovirus B3) ATCC VR-3 4993.0 TCID50/ml 20/20
C35* ATCC VR-94 0.2 TCID50/ml 19/20
–
Greer ATCC VR-92 7.3 TCID50/ml 20/20
–* ZeptoMetrix
C 243 ATCC VR-93 2.3 TCID50/ml 20/20
–* ZeptoMetrix
M-25 ATCC VR-1378 0.5 TCID50/ml 20/20
–* ZeptoMetrix
A2* ATCC VR-1540 12.0 PFU/ml 20/20
Long* ATCC VR-26 33.0 PFU/ml 20/20
18537* ATCC VR-1580 0.03 PFU/ml 20/20
CH93(18)-18 ZeptoMetrix
Peru6-2003 (type B2)* ZeptoMetrix
hMPV-16, IA10-2003 (A1) ZeptoMetrix
hMPV-20, IA14-2003 (A2)*
hMPV-3, Peru2-2002 (B1)*
RI-67 (Adenovirus E4)* ATCC VR-1572 15.8 TCID50/ml 20/20
Adenoid 75 (Adenovirus C5)* ATCC VR-5 7331.0 TCID50/ml 20/20
Adenoid 71 (Adenovirus C1)* ATCC VR-1 69.5 TCID50/ml 20/20
Adenoid 6 (Adenovirus C2)* ATCC VR-846 28.1 TCID50/ml 20/20
Tonsil 99 (Adenovirus C6)* ATCC VR-6 88.8 TCID50/ml 20/20
ZeptoMetrix
0810014CFHI
0810015CFHI
0810016CFHI
0810060BCFHI
0810040CFHI
0810159CFHI
0810161CFHI
ZeptoMetrix,
0810163CFHI
ZeptoMetrix,
0810156CFHI
0.2 TCID50/ml 19/20
1.3 TCID50/ml 19/20
11.5 TCID50/ml 20/20
9.5 TCID50/ml 20/20
0.4 TCID50/ml 20/20
0.01 TCID50/ml 19/20
0.5 TCID50/ml 20/20
0.4 TCID50/ml 19/20
1479.9 TCID50/ml 19/20
(continued on next page)

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
83
Table 18 (continued)
Pathogen Strain Source Concentration Detection rate
Enterovirus /US/IL/14-18952 (Enterovirus
Rhinovirus 1059 (Rhinovirus B14)* ATCC VR-284 8.9 TCID50/ml 20/20
Mycoplasma
pneumoniae
Chlamydia
pneumoniae
Bordetella pertussis
* The LoD has been obtained in simulated matrix.
D68)
Echovirus 6* ATCC VR-241 0.9 TCID50/ml 19/20
HGP (Rhinovirus A2) ATCC VR-482 8.9 TCID50/ml 19/20
11757 (Rhinovirus C16)* ATCC VR-283 50.0 TCID50/ml 20/20
Type 1A* ATCC VR-1559 8.9 TCID50/ml 20/20
M129-B7 (type 1)* ATCC 29342 0.1 CCU/ml 20/20
PI 1428 ATCC 29085 1.0 CCU/ml 20/20
TW183 ATCC VR-2282 14.2 IFU/ml 20/20
CWL-029* ATCC VR-1310 120.0 IFU/ml 19/20
I028 ATCC BAA-
18323* ATCC 9797 2.6 CFU/ml 19/20
ATCC VR-1824 8.9 TCID50/ml 19/20
0.3 CFU/ml 20/20
2707
Exclusivity (Cross-reactivity and Exclusivity)
The analytical specificity study was carried out by
in silico
analysis and
in vitro
testing to assess the
cross-reactivity and exclusivity of the QIAstat-Dx Respiratory Panel. On-panel organisms were tested
to assess the potential for intra-panel cross-reactivity and off-panel organisms were tested to evaluate
panel exclusivity. The off-panel organisms selected were clinically relevant organisms (colonizing
the upper respiratory tract or causing respiratory symptoms), common skin flora or laboratory
contaminants, or microorganisms for which much of the population may have been infected. The
on-panel organisms tested are shown in Table 20 (next page).
Samples were prepared by spiking potential cross-reactive organisms into simulated
nasopharyngeal swab sample matrix at the highest concentration possible based on the
5
organism stock – at least 10
TCID50/ml for viral targets and 106 CFU/ml for bacterial and
fungal targets. These concentrations represent levels approximately 800–1,000,000-fold
higher than the LoD of the QIAstat-Dx Respiratory Panel.

84
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Bordetella pertussis
detection in the QIAstat-Dx Respiratory SARS-CoV-2 Panel targets the
IS481 region. As a consequence, a certain level of cross-reactivity with off-panel
species and
Bordetella pertussis
observed when high concentrations of
was predicted by preliminary sequence analysis and was
Bordetella holmesii
were tested. In concordance with
the CDC guidelines for assays that use the IS481 as a target region, when using QIAstat-Dx
Respiratory SARS-CoV-2 Panel if the C
value for
T
Bordetella pertussis
is CT>29, a confirmatory
specificity test is recommended.
Table 20. List of Analytical Specificity pathogens tested
Type Pathogen
Mycoplasma pneumoniae
On-panel bacteria
Off-panel bacteria
Bordetella pertussis
Chlamydia pneumoniae
Acinetobacter calcoaceticus
Bordetella avium
Bordetella bronchiseptica
Bordetella hinzii
Bordetella holmesii
Bordetella parapertussis
Chlamydia trachomatis
Corynebacterium diphteriae
Enterobacter aerogenes
Escherichia coli
Haemophilus influenzae
Klebsiella oxytoca
Klebsiella pneumoniae
Lactobacillus acidophilus
Lactobacillus plantarum
Legionella bozemanii
Legionella dumofii
Legionella feeleii
Legionella longbeacheae
Legionella micdadei
Legionella pneumophila
Moraxella catarrhalis
(O157)
(continued on next page)
Bordetella

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
85
Table 20 (continued)
Type Pathogen
Mycobacterium tuberculosis*
Mycoplasma genitalium
Mycoplasma hominis
Mycoplasma orale
Neisseria elongata
Neisseria gonorrhoeae
Neisseria meningitidis
Proteus mirabilis
Off-panel bacteria (continued)
On-panel viruses
Pseudomonas aeruginosa
Serratia marcescens
Staphylococcus aureus
Staphylococcus epidermidis
Stenotrophomonas maltophilia
Streptococcus agalactiae
Streptococcus pneumoniae
Streptococcus pyogenes
Streptococcus salivarus
Ureaplasma urealyticum
Influenza A H1N1
Influenza A H3N2
Influenza A H1N1/pdm09
Influenza B
Coronavirus 229E
Coronavirus OC43
Coronavirus NL63
Coronavirus HKU1
Parainfluenza virus 1
Parainfluenza virus 2
Parainfluenza virus 3
Parainfluenza virus 4a
RSV A
hMPV A
Adenovirus C
Adenovirus B
†
(continued on next page)

86
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Table 20 (continued)
Type Pathogen
On-panel viruses (continued)
Off-panel viruses
Off-panel fungi
*
Mycobacterium tuberculosis
†
Coronavirus HKU1 clinical specimen tested.
‡
Bocavirus Type 1 clinical specimens tested.
§
Middle East Respiratory Syndrome Coronavirus synthetic RNA tested.
genomic DNA tested.
Enterovirus
Rhinovirus
Bocavirus‡
Cytomegalovirus
Epstein-Barr Virus
Herpes Simplex Virus 1
Herpes Simplex Virus 2
Measles Virus
Middle East Respiratory Syndrome Coronavirus§
Mumps
Aspergillus flavus
Aspergillus fumigatus
Candida albicans
Cryptococcus neoformans
Inclusivity (Analytical Reactivity)
Analytical reactivity (Inclusivity) was evaluated with a collection of 127 respiratory pathogen
isolates/strains that were selected based on clinical relevance and temporal/geographical
diversity. Based on wet testing and
Panel primers and probes are specific and inclusive for clinically prevalent and relevant strains
in silico
for each pathogen. Wet testing has been done with the strains listed in Table 20. Every strain
has been tested in triplicate with a 100% detection rate for concentrations listed.
Table 21 (following pages) provides details of the respiratory pathogens tested in this study.
analysis, the QIAstat-Dx Respiratory SARS-CoV-2

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
87
Table 21. List of Analytical Reactivity pathogens tested
Pathogen
Influenza A
Subtype/
serotype
H1N1
H3N2
Strain Source
A/PR/8/34 ATCC VR-1469 3x Influenza A H1
A/New Jersey/8/76* ATCC VR-897 1x Influenza A H1
A/Brisbane/59/07*
A/New
Caledonia/20/99*
A/Denver/1/57 ATCC VR-546 0.1x Influenza A H1
A/Weiss/43 ATCC VR-96 0.1x Influenza A H1
A/Fort Monmouth/1/1947 ATCC VR-1754 0.1x Influenza A H1
A/WS/33 ATCC VR-1520 0.1x Influenza A H1
A/Swine/Iowa/15/1930 ATCC VR-333 1x Influenza A H1
A/Mal/302/54 ATCC VR-98 1x Influenza A H1
A/Virginia/ATCC6/2012* ATCC VR-1811 1x Influenza A H3
A/Wisconsin/67/2005* ZeptoMetrix
A/Port Chalmers/1/73* ATCC VR-810 1x Influenza A H3
A/Victoria/3/75 ATCC VR-822 1x Influenza A H3
A/Aichi/2/68 ATCC VR-1680 10x Influenza A H3
A/Hong Kong/8/68 ATCC VR-1679 10x Influenza A H3
A/Alice (recombinant,
carries A/England/42/72)
MRC-2 (recombinant
A/England/42/72 and
A/PR/8/34 strains)
A/Switzerland/
9715293/2013
A/Wisconsin/15/2009 ATCC VR-1882 1x Influenza A H3
ZeptoMetrix
0810244CFHI
ZeptoMetrix
0810036CFHI
0810252CFHI
ATCC VR-776 10x Influenza A H3
ATCC VR-777 100x Influenza A H3
ATCC VR-1837 1x Influenza A H3
x LoD
Detected
1x Influenza A H1
0.3x Influenza A H1
1x Influenza A H3
Result
* Strain tested during LoD verification study.
Note: Influenza A/Brisbane/59/07 (ZeptoMetrix, 0810244CFHI) taken as reference strain to calculate the x-fold LoD
detected.
(continued on next page)

88
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Table 21 (continued)
Pathogen
Influenza A
Subtype/
serotype
H1N1
(pandemic)
H2N2
H5N3
Strain Source
A/Virginia/ATCC1/2009* ATCC VR-1736 1x Influenza A
A/SwineNY/03/2009* ZeptoMetrix
Canada/6294/09 ZeptoMetrix
Mexico/4108/09 ZeptoMetrix
Netherlands/2629/2009
A/Virginia/ATCC2/2009 ATCC VR-1737 0.1x
A/Virginia/ATCC3/2009 ATCC VR-1738 100x
Swine NY/01/2009
Swine NY/02/2009
A/California/07/2009
NYMC X-179A
Japan/305/1957 (nucleic
acid)
Korea/426/1968xPuerto
Rico/8/1934 [isolate
ID/source=recombinant]
(nucleic acid)
A/Duck/Singapore/645/
1997 [isolate
ID/source=avian]
0810249CFHI
0810109CFJHI
0810166CFHI
BEI Resources NR19823
ZeptoMetrix
0810248CFHI
ZeptoMetrix
0810109CFNHI
ATCC VR-1884 0.1x
BEI Resources NR2775
BEI Resources NR9679
BEI Resources NR9682
x LoD
Detected
1x Influenza A
3x Influenza A
0.1x Influenza A
0.3x
0.3x
10x
1x Influenza A (no
0.3x Influenza A (no
1x
Result
H1N1/pdm09
H1N1/pdm09
H1N1/pdm09
H1N1/pdm09
Influenza A
H1N1/pdm09
Influenza A
H1N1/pdm09
Influenza A
H1N1/pdm09
Influenza A
H1N1/pdm09
Influenza A
H1N1/pdm09
Influenza A
H1N1/pdm09
subtype)
subtype)
Influenza A (no
subtype)
* Strain tested during LoD verification study.
(continued on next page)

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
89
Table 21 (continued)
Pathogen
Subtype/
serotype
Strain Source
H10N7 Chicken/Germany/N/49
[isolate ID/source=avian]
BEI Resources NR2765
x LoD
Detected
Result
10x Influenza A (no
subtype)
(nucleic acid)
Influenza A
H1N2 Recombinant Kilbourne
F63, A/NWS/1934 (HA) x
A/Rockefeller
BEI Resources NR9677
100x Influenza A H1
Institute/5/1957 (NA)
[isolate
ID/source=recombinant]
(nucleic acids)
B/Virginia/ATCC5/2012* ATCC VR-1807 1x Influenza B
B/FL/04/06* ATCC VR-1804 1x Influenza B
B/Taiwan/2/62* ATCC VR-295 0.3x Influenza B
B/Allen/45 ATCC VR-102 Not
detected
B/Hong Kong/5/72 ATCC VR-823 Not
detected
†
†
Negative
Negative
B/Maryland/1/59 ATCC VR-296 0.1x Influenza B
Influenza B
Not
available
B/GL/1739/54 ATCC VR-103 1x Influenza B
B/Wisconsin/1/2010 ATCC VR-1883 0.1x Influenza B
B/Massachusetts/2/2012 ATCC VR-1813 3x Influenza B
‡
Influenza B or
Negative
B/Florida/02/06 ZeptoMetrix
0810037CFHI
Impaired
detectability
B/Brisbane/60/2008 BEI Resources NR-
42005
B/Malaysia/2506/2004 BEI Resources NR-
0.1x Influenza B
0.3x Influenza B
9723
* Strain tested during LoD verification study.
†
Both strains are derivative from B/Lee/40 ancestral lineage and, according to
to be detected by the QIAstat-Dx Respiratory Panel.
‡
In silico
analysis showed that this strain should be detected by QIAstat-Dx Respiratory Panel.
in silico
analysis, they were predicted
(continued on next page)

90
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Table 21 (continued)
Pathogen
Coronavirus
229E
Coronavirus
OC43
Coronavirus
NL63
Coronavirus
HKU1
Subtype/
serotype
Not
available
Not
available
Not
available
Not
available
Not
available
Strain Source
Not available* ZeptoMetrix
Not available* ATCC VR-740 0.3x Coronavirus
Not available* ATCC VR-1558 1x Coronavirus
Not available* ZeptoMetrix
Not available*
Not available
Not available*
Not available
Not available QIAGEN
Not available
0810229CFHI
0810024CFHI
ZeptoMetrix
0810228CFHI
BEI Resources NR470
ZeptoMetrix
NATRVP-IDI
QIAGEN
Barcelona (STATDx) Clinical sample
S510†
Barcelona (STATDx) Clinical sample
S501†
QIAGEN
Barcelona (STATDx) Clinical sample
496†
x LoD
Detected
1x Coronavirus
1x Coronavirus
1x
1x
1x
0.3x
1x Coronavirus
1x
Result
229E
229E
OC43
OC43
Coronavirus
NL63
Coronavirus
NL63
Coronavirus
HKU1
Coronavirus
HKU1
HKU1
Coronavirus
HKU1
* Strain tested during LoD verification study.
†
Stock titer not available according to manufacturer.
(continued on next page)

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
91
Table 21 (continued)
Pathogen
Parainfluenza
virus 1
Subtype/
serotype
Not
available
Strain Source
Not available* ZeptoMetrix
0810014CFHI
C35* ATCC VR-94 1x Parainfluenza
Not available ZeptoMetrix
NATRVP-IDI
†
x LoD
Detected
1x Parainfluenza
10x Parainfluenza
Result
virus 1
virus 1
virus 1
Greer* ATCC VR-92 1x Parainfluenza
virus 2
Parainfluenza
virus 2
Parainfluenza
virus 3
Not
available
Not
available
Not available*
ZeptoMetrix
0.3x
0810015CFHI
Not available
ZeptoMetrix
0.1x
0810504CFHI
Not available*
ZeptoMetrix
1x
0810016CFHI
C 243* ATCC VR-93 1x
Not available*
ZeptoMetrix
NATRVP-IDI
†
0.1x
M-25* ATCC VR-1378 1x
Parainfluenza
virus 2
Parainfluenza
virus 2
Parainfluenza
virus 3
Parainfluenza
virus 3
Parainfluenza
virus 3
Parainfluenza
virus 4
Parainfluenza
virus 4
Not
available
Not available*
ZeptoMetrix
0.3x
0810060BCFHI
Not available
ZeptoMetrix
0.1x
0810060CFHI
CH 19503 ATCC VR-1377 0.3x
Parainfluenza
virus 4
Parainfluenza
virus 4
Parainfluenza
virus 4
* Strain tested during LoD verification study.
†
Stock titer not available according to manufacturer.
(continued on next page)

92
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Table 21 (continued)
Pathogen
Respiratory
Syncytial
Virus A+B
Human Metapneumovirus
Subtype/
serotype
Not
available
Not
available
Strain Source
18537* ATCC VR-1580 1x Respiratory Syncytial
A2* ATCC VR-1540 0.3x Respiratory Syncytial
Long* ATCC VR-26 1x Respiratory Syncytial
CH93(18)-18* ZeptoMetrix
Not available
B WV/14617/85 ATCC VR-1400 1x
IA10-2003*
IA14-2003*
Peru2-2002*
Peru6-2003*
IA3-2002
IA27-2004
Peru3-2003
IA18-2003
Peru1-2002 ZeptoMetrix
0810040CFHI
ZeptoMetrix
0810040ACFHI
ZeptoMetrix
0810161CFHI
ZeptoMetrix
0810163CFHI
ZeptoMetrix
0810156CFHI
ZeptoMetrix
0810159CFHI
ZeptoMetrix
0810160CFHI
ZeptoMetrix
0810164CFHI
ZeptoMetrix
0810158CFHI
ZeptoMetrix
0810162CFHI
0810157CFHI
x LoD
Detected
1x Respiratory Syncytial
0.1x
1x
1x
1x
1x
3x
1x
1x
1x
10x Human Meta-
Result
virus A+B
virus A+B
virus A+B
virus A+B
Respiratory Syncytial
virus A+B
Respiratory Syncytial
virus A+B
Human Metapneumovirus A+B
Human Metapneumovirus A+B
Human Metapneumovirus A+B
Human Metapneumovirus A+B
Human
Metapneumovirus A+B
Human Metapneumovirus A+B
Human Metapneumovirus A+B
Human Metapneumovirus A+B
pneumovirus A+B
* Strain tested during LoD verification study.
(continued on next page)

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
93
Table 21 (continued)
Pathogen
Adenovirus
* Strain tested during LoD verification study.
Subtype/
serotype
Not
available
Strain Source
Tonsil 99* ATCC VR-6 1x Adenovirus
GB* ATCC VR-3 0.3x Adenovirus
Adenoid 71* ATCC VR-1 1x Adenovirus
Adenoid 6* ATCC VR-846 0.3x Adenovirus
Adenoid 75* ATCC VR-5 0.3x Adenovirus
RI-67* ATCC VR-1572 0.3x Adenovirus
Huie ATCC VR-863 0.3x Adenovirus
Gomen ATCC VR-7 0.1x Adenovirus
Slobitski ATCC VR-12 10x Adenovirus
AV-1645 [128] ATCC VR-256 0.3x Adenovirus
Compton ATCC VR-716 0.3x Adenovirus
Holden ATCC VR-718 0.3x Adenovirus
Trim ATCC VR-1815 0.3x Adenovirus
Dugan ATCC VR-931 0.1x Adenovirus
Tak (73-3544) ATCC VR-930 3x Adenovirus
x LoD
Detected
Result
(continued on next page)

94
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Table 21 (continued)
Pathogen
Enterovirus
Rhinovirus
Subtype/
serotype
Not
available
Not
available
Strain Source
/US/IL/1418952*
D-1 (Cox)* ATCC VR-241 0.3x Rhinovirus/Enterovirus
H ATCC VR-1432 1x Rhinovirus/Enterovirus
M.K. (Kowalik) ATCC VR-168 10x Rhinovirus/Enterovirus
Gregory ATCC VR-41 10x Rhinovirus/Enterovirus
Bastianni ATCC VR-1660 1x Rhinovirus/Enterovirus
Griggs ATCC VR-1311 0.3x Rhinovirus/Enterovirus
Conn-5 ATCC VR-28 0.3x Rhinovirus/Enterovirus
Ohio-1 ATCC VR-29 3x Rhinovirus/Enterovirus
Nancy ATCC VR-30 0.3x Rhinovirus/Enterovirus
CHHE-29 ATCC VR-47 10x Rhinovirus/Enterovirus
Kuykendall [V-024001-012]
1059* ATCC VR-284 1x Rhinovirus/Enterovirus
2060* ATCC VR-1559 0.1x Rhinovirus/Enterovirus
HGP* ATCC VR-482 1x Rhinovirus/Enterovirus
11757* ATCC VR-283 0.3x Rhinovirus/Enterovirus
FEB ATCC VR-483 1x Rhinovirus/Enterovirus
33342 ATCC VR-1663 3x Rhinovirus/Enterovirus
ATCC VR-1824 1x Rhinovirus/Enterovirus
ATCC VR-850 10x Rhinovirus/Enterovirus
x LoD
Detected
Result
* Strain tested during LoD verification study.
(continued on next page)

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
95
Table 21 (continued)
Pathogen
M.
pneumoniae
B. pertussis
C.
pneumoniae
* Strain tested during LoD verification study.
†
Stock titer not available according to manufacturer.
Subtype/
serotype
Not
available
Not
available
Not
available
Interfering substances
Strain Source
PI 1428* ATCC 29085 1x
M129-B7* ATCC 29342 1x
FH strain of Eaton Agent
[NCTC 10119]
I028* ATCC BAA-2707 1x
19323* ATCC 9797 1x
10-536 ATCC 10380† 0.3x
TW183* ATCC VR-2282 1x
CWL-029* ATCC VR-1310 1x
AR-39 ATCC 53592 0.3x
ATCC 15531 0.1x
x LoD
Detected
Result
Mycoplasma
pneumoniae
Mycoplasma
pneumoniae
Mycoplasma
pneumoniae
Bordetella
pertussis
Bordetella
pertussis
Bordetella
pertussis
Chlamydophila
pneumoniae
Chlamydophila
pneumoniae
Chlamydophila
pneumoniae
The effect of potentially interfering substances on the detectability of the QIAstat-Dx Respiratory
Panel organisms was evaluated. Thirty (30) potentially interfering substances were added to
contrived samples at a level predicted to be above the concentration of the substance likely to
be found in an authentic NPS specimen. The contrived samples (also referred to as combined
samples) were each comprised of a mix of organisms tested at a concentration of 5x LoD.
Endogenous substances such as whole blood, human genomic DNA and several pathogens
were tested alongside exogenous substances like antibiotics, nasal sprays and different
workflow contaminants.

96
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
The combined samples were tested with and without addition of an inhibitory substance
allowing direct sample-to-sample comparison. Combined samples not spiked with any test
substance served as a positive control. Additionally, for substances that may contain genetic
material (such as blood, mucin, DNA and microorganisms), negative specimens (blank sNPS
sample matrix with no organism mix) were spiked with only the test substance to evaluate the
potential for false positive results due to the test substance itself.
Combined samples not spiked with any test substance served as a positive control and blank
sNPS sample matrix with no organism mix as negative controls.
All pathogen-containing samples without spiked interferent generated positive signals for all
pathogens present in the respective combined sample. Negative signals were obtained for all
pathogens not present in the same sample but detected by the QIAstat-Dx Respiratory Panel.
None of the substances tested showed inhibition, except for the nasal influenza vaccines. This
was due to the fact that the selection of substances concentration was higher than the
concentrations expected to be present in a sample. In addition, nasal influenza vaccines
®
(Fluenz Tetra and FluMist
) were predicted to be reactive with the QIAstat-Dx Respiratory Panel
Influenza A (subtype) and Influenza B assays. Final dilution without observable interfering
effect was 0.000001% v/v for both vaccines.
No impact on performance is expected when clinical liquid samples are examined in the
presence of the substances tested.
Clinically relevant co-infections testing demonstrated that when at least two QIAstat-Dx
Respiratory Panel pathogens of different concentrations are simultaneously present in one
sample all targets can be detected by the assay.
The results of interfering substance testing are provided in Table 22 (next page).

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
97
Table 22. Final highest concentration without observable inhibitory effect
Substance tested Concentration tested Results
Endogenous substances
Human genomic DNA 200 ng/µL 20 ng/µl No Interference
Human blood (+NaCitrate) 1% v/v No Interference
Mucin from bovine submaxillary 1% v/v No Interference
Competitive microorganisms
Staphylococcus aureus
Neisseria meningitidis
Corynebacterium diphtheriae
Human Cytomegalovirus 1.00E+05 TCID50/ml No Interference
Exogenous substances
Tobramycin 0.6 mg/ml No Interference
Mupirocin 2% w/v No Interference
Saline nasal spray with preservatives 1% v/v No Interference
Afrin®, severe congestion nasal spray (Oxymetazoline
HCl)
Analgesic ointment (Vicks® VapoRub®) 1% w/v No Interference
Petroleum Jelly (Vaseline®) 1% w/v No Interference
FluMist nasal influenza vaccine 0.00001% v/v Interference
FluMist nasal influenza vaccine 0.000001% v/v No Interference
Fluenz Tetra nasal influenza vaccine 0.00001% v/v Interference
Fluenz Tetra nasal influenza vaccine 0.000001% v/v No Interference
Disinfecting/cleaning substances
Disinfecting wipes ½ inches2/1 ml UTM No Interference
DNAZap™ 1% v/v No Interference
RNaseOUT™ 1% v/v No Interference
Bleach 5% v/v No Interference
Ethanol 5% v/v No Interference
1.00E+06 CFU/ml No Interference
5.00E+04 CFU/ml No Interference
5.00E+03 CFU/ml No Interference
1% v/v No Interference
(continued on next page)

98
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Table 22 (continued)
Substance tested Concentration tested Results
Specimen collection materials
Swab Copan 168C 1 swab/1 ml UTM No Interference
Swab Copan FloQ 1 swab/1 ml UTM No Interference
Swab Copan 175KS01 1 swab/1 ml UTM No Interference
Swab Puritan 25-801 A 50 1 swab/1 ml UTM No Interference
VTM Sigma Virocult
VTM Remel® M4RT
VTM Remel M4
VTM Remel M5 100% No Interference
VTM Remel M6 100% No Interference
BD Universal Viral Transport 100% No Interference
100% No Interference
100% No Interference
100% No Interference
Reproducibility
Reproducibility testing of contrived samples was performed at three test sites including two
external sites (LACNY [Laboratory Alliance of Central New York] and INDIANA [Indiana
University]) and one internal site (STAT). The study incorporated a range of potential variation
introduced by sites, days, replicates, cartridge lots, operators, and QIAstat-Dx analyzers. For
each site, testing was performed across 5 days with 4 replicates per day (leading to a total of
20 replicates per target, concentration and site), a minimum of 2 different QIAstat-Dx
Analyzers per site, and at least 2 operators on each testing day.
A total of 12 sample mixes were prepared with at least 3 replicates tested per sample mix.
Each pathogen was spiked into HeLa in UTM combined samples in a final concentration of
0.1x LoD, 1x LoD or 3x LoD, respectively. A summary of results for each analyte is provided
in Table 23 (next page).
Table 23 summarizes the results for 0.1x LoD concentration where it is observed that the
detection rate for 24 of the 24 targets was <95% and therefore the acceptance criteria is met.

QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
99
Reproducibility and repeatability will impact the SARS-CoV-2 target in the same manner as
other target organisms verified in the QIAstat-Dx Respiratory Panel.
Table 23. Detection rate per target at 0.1x LoD concentration for each site of reproducibility study and 2-sided 95%
Confidence Interval by target
Target
(0.1x LoD) Site
STAT 10/20 50.0% 29.9–70.1%
Adenovirus
(ATCC VR-3)
B. pertussis
(BAA-2707)
C. pneumoniae
(ATCC VR-
2282)
Coronavirus
229E (ATCC VR-
740)
Coronavirus
HKU1 (NATRVPIDI)
Coronavirus
NL63
(0810228CFHI)
LACNY 9/19 47.4% 27.3–68.3%
INDIANA 10/19 52.6% 31.7–72.7%
All sites (overall) 29/58 50.0% 37.5–62.5%
STAT 9/20 45.0% 25.8–65.8%
LACNY 7/19 36.8% 19.1–59.0%
INDIANA 9/20 45.0% 25.8–65.8%
All sites (overall) 25/59 42.4% 30.6–55.1%
STAT 11/20 55.0% 34.2–74.2%
LACNY 11/19 57.9% 36.3–76.9%
INDIANA 14/20 70.0% 48.1–85.5%
All sites (overall) 36/59 61.0% 48.3–72.4%
STAT 9/20 45.0% 25.8–65.8%
LACNY 12/19 63.2% 41.0–80.9%
INDIANA 5/20 25.0% 11.2–46.9%
All sites (overall) 26/59 44.1% 32.2–56.7%
STAT 17/20 85.0% 64.0–94.8%
LACNY 10/19 52.6% 31.7–72.7%
INDIANA 9/20 45.0% 25.8–65.8%
All sites (overall) 36/59 61.0% 48.3–72.4%
STAT 13/20 65.0% 43.3–81.9%
LACNY 12/19 63.2% 41.0–80.9%
INDIANA 14/19 73.7% 51.2–88.2%
All sites (overall) 39/58 67.2% 54.4–77.9%
Detection rate
(# positive)
% detection rate (#
positive)
95% Confidence
Interval
(continued on next page)

100
QIAstat-Dx Respiratory SARS-CoV-2 Panel Instructions for Use (Handbook) 09/2020
Table 23 (continued)
Target
(0.1x LoD)
Coronavirus
OC43 (ATCC
VR-1558)
Enterovirus
(ATCC VR-1824)
Human
Metapneumovirus
(0810161CF)
Influenza A
(0810249CFHI)
Influenza A
(ATCC VR-810)
Influenza A
(ATCC VR-897)
Site
STAT 13/20 65.0% 43.3–81.9%
LACNY 15/20 75.0%
INDIANA 15/20 75.0% 53.1–88.8%
All sites (overall) 43/60 71.7% 59.2–81.5%
STAT 8/20 40.0% 21.9–61.3%
LACNY 6/19 31.6% 15.4–54.0%
INDIANA 7/20 35.0% 18.1–56.7%
All sites (overall) 21/59 35.6% 24.6–48.3%
STAT 6/20 30.0% 14.5–51.9%
LACNY 9/19 47.4% 27.3–68.3%
INDIANA 9/20 45.0% 25.8–65.8%
All sites (overall) 24/59 40.7% 29.1–53.4%
STAT 19/20 95.0% 76.4–99.1%
LACNY 18/20 90.0% 69.9–97.2%
INDIANA 20/20 100% 83.9–100%
All sites (overall) 57/60 95.0% 86.3–98.3%
STAT 10/20 50.0% 29.9–70.1%
LACNY 9/19 47.4% 27.3–68.3%
INDIANA 16/19 84.2% 62.4–94.5%
All sites (overall) 35/58 60.3% 47.5–71.9%
STAT 14/20 70.0% 48.1–85.5%
LACNY 9/19 47.4% 27.3–68.3%
INDIANA 12/20 60.0% 38.7–78.1%
All sites (overall) 35/59 59.3% 46.6–70.9%
Detection rate
(# positive)
% detection rate (#
positive)
95% Confidence
Interval
53.1–88.8%
(continued on next page)
 Loading...
Loading...